
(DahlL,etal. SciAdv.2023)
The G protein-coupled receptor (GPCR) superfamily is the largest family of human membrane proteins and drug targets, comprising approximately 750 membrane proteins. Their function primarily relies on their ability to change shape and transition between different conformations. They mediate numerous signaling pathways involved in cellular physiology and intercellular communication. Disruption of GPCR function or signal transduction is implicated in the pathophysiology of many disease states.
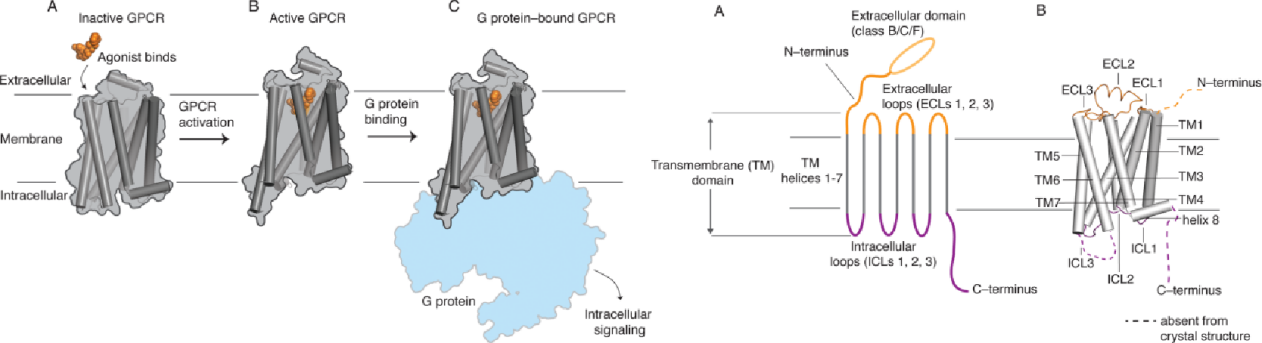
(LatorracaNR,etal. ChemRev.2017)
Therefore, G protein-coupled receptors (GPCRs) are also targets for approximately 30% of approved therapeutic drugs. Although GPCRs represent one of the most targeted protein families in drug development, the dynamic nature of these membrane proteins poses challenges and limitations for the development of specific antibodies.
In May 2023, a research paper titled "Multiplexed selectivity screening of anti-GPCR antibodies" was published online in Advanced Science. The study developed a multiplex immunoassay method to test 407 anti-GPCR antibodies against a customized library of 215 GPCRs. Based on the data results, it provides insights into the immunogenicity of GPCR family proteins and the design and application of anti-GPCR antibodies as tool reagents and therapeutic agents.

Establishment of a Multiplexed Detection Method for Anti-GPCR Antibodies:
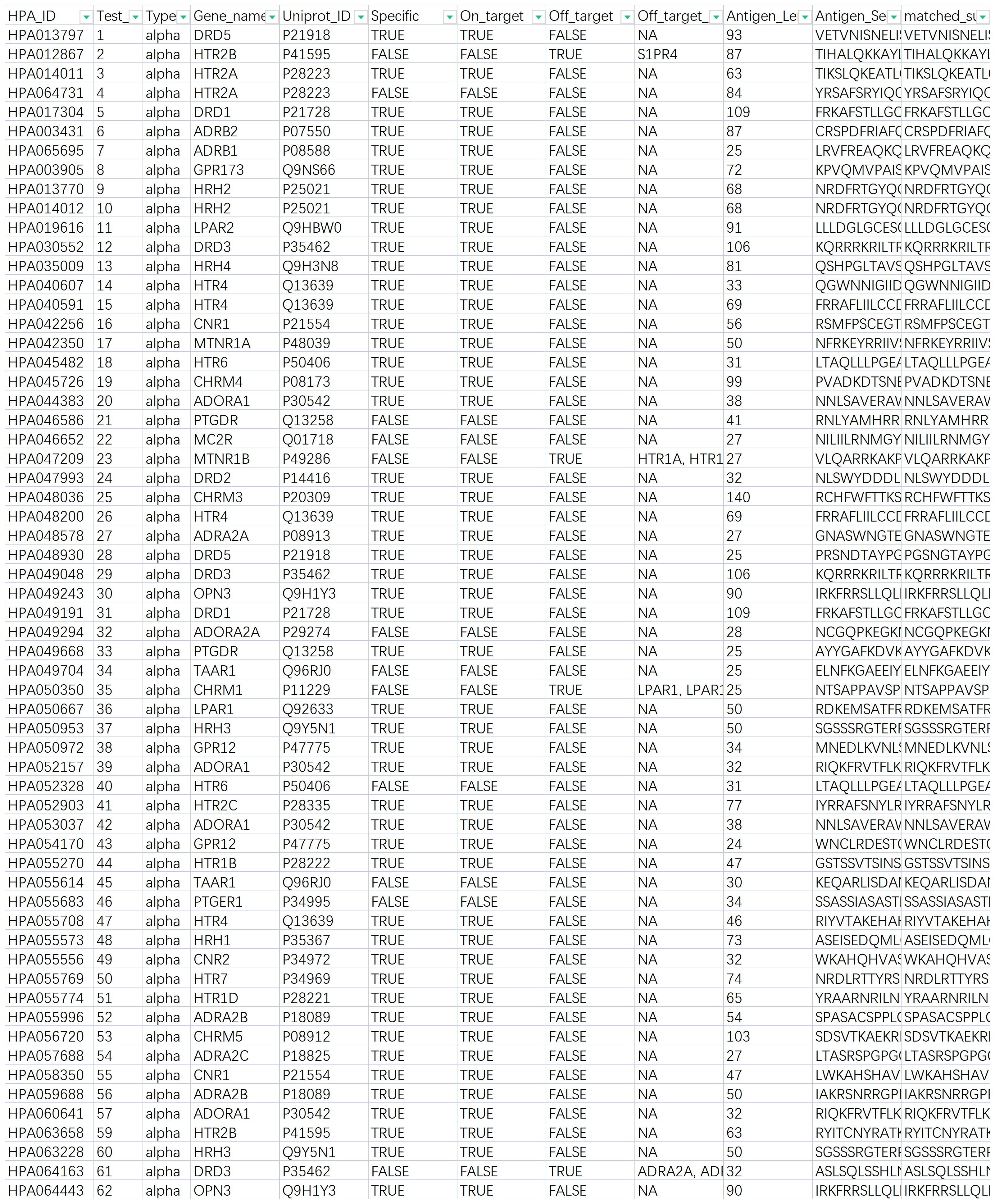
Researchers combined parallel expression of GPCRs with multiplexed protein capture analysis, using engineered tags to detect immunocaptured proteins and determine the relative quantities of GPCRs in solution. The data were then integrated into a network-based interface to explore the selectivity of each antibody for GPCR proteins of the 407 anti-GPCR antibodies tested, the majority were sourced from the Human Protein Atlas (HPA) program. The GPCR library was created by ectopic expression of 215 dual-site labeled GPCRs in 293F cells, each GPCR protein including an N-terminal Flag tag and a C-terminal 1D4 tag. Dodecylmaltoside (DM) was used as a detergent for washing, generating detergent micelles containing heterogeneous mixtures. Through multiplexed protein capture assays, the researchers determined which antibodies were selective in recognizing the intended GPCR targets.

(Experimental workflow and data analysis)
Discovery of Selective GPCR Antibodies:
In the selective binding screening of 407 antibodies against 205 GPCRs (although there were 215 GPCRs in the library, antibodies selected for 10 GPCRs did not pass QC testing and were only used as off-target candidates), 61% (248 out of 407) of the tested antibodies recognized their intended GPCRs. Of the remaining 159 antibodies, 9% (15 out of 159) enriched the intended targets along with at least one off-target, while an additional 20% (31 out of 159) of the remaining antibodies only bound to an unintended target, and 71% (113 out of 159) did not enrich any targets beyond the cutoff. The 248 highly selective antibodies corresponded to 154 unique GPCR targets.
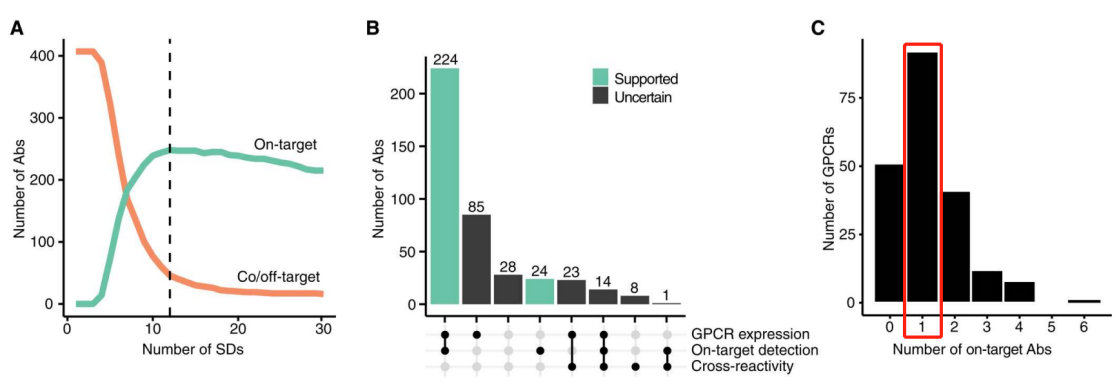
(Antibody Selectivity Thresholds and Summary Statistics)
Analysis of Different GPCR Sites:
Among the 205 GPCRs targeted by antibodies, 116 (57%) were targeted by two or more pairs of antibodies, allowing researchers to compare binding features with GPCRs and present different sites for selective and non-selective recognition of GPCRs.
For the rhodopsin family, the α-subfamily member adrenergic receptor α2B (ADRA2B ) was selectively recognized by six of six antibodies, which were raised against three different epitope regions, all located in the intracellular loop 3 of ADRA2B. (ICL3); the β-subfamily member gastrin-releasing peptide receptor was selectively recognized by five of the six antibodies, raised against two different epitope regions, one within the extracellular domain (ECD) and one within extracellular loop 2 (ECL2); the δ-subfamily member CCR7 was selectively recognized by only one of the five antibodies, HPA060045, despite using three different antigens: one in the ECD, one in ECL2, and one corresponding to the intracellular C-terminal tail of the receptor (CCR7 is expressed at lower levels than most receptors of the δ-subfamily and was therefore classified as “uncertain,” which may indicate that only antibodies with sufficiently high affinity for CCR7 can detect even lower levels of this GPCR); the γ-subfamily member follicle-stimulating hormone receptor (FSHR) was selectively recognized by three, all with three different epitope regions within the large ECD of FSHR.
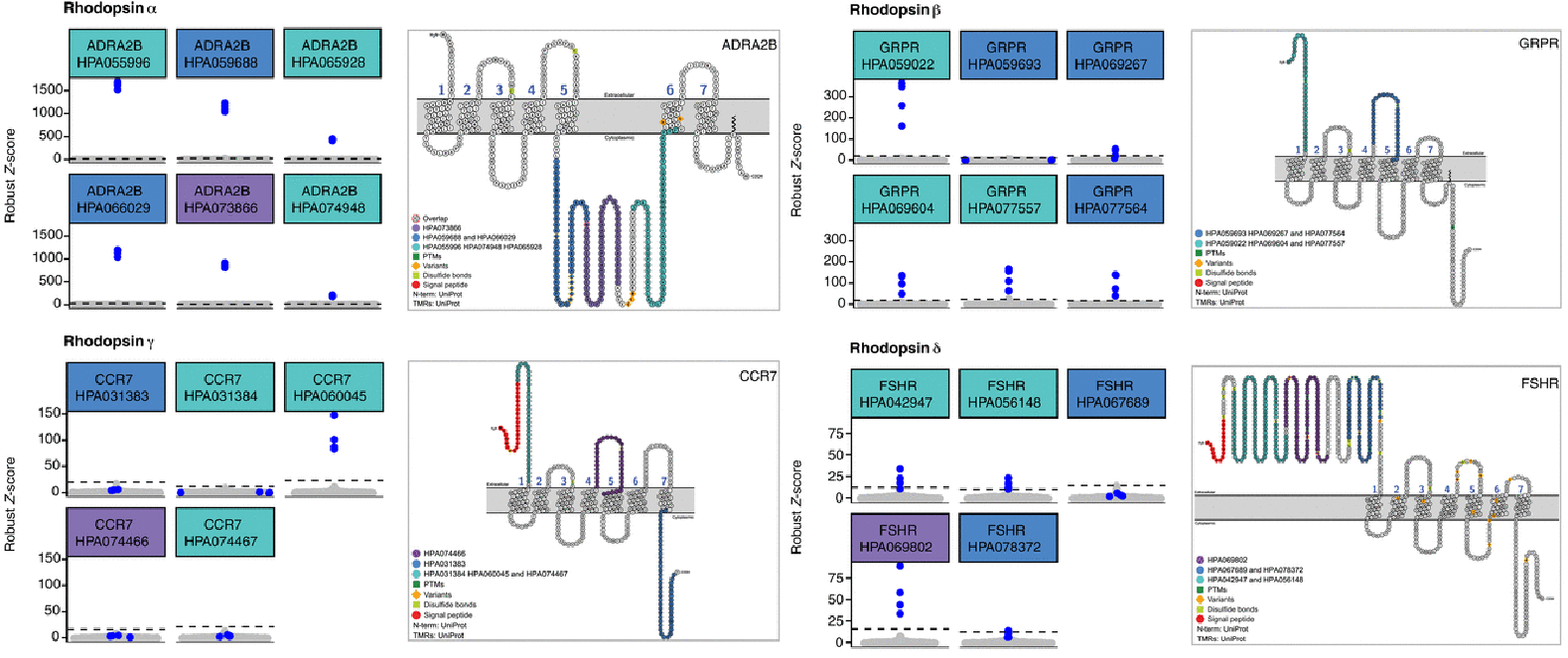
(Rhodopsin Family Protein Recognition Site Map)
Within the Glutamate Family, the GPCR class C group 5 member D (GPRC5D) was selectively recognized by three of four antibodies. The failed antibody, HPA047203, was labeled to recognize the ECL2 region, while selective antibodies targeted its ECD (HPA071909) or the intracellular C-terminal tail (HPA064241 and HPA071739). HPA071739 cross-reacted with the related receptor GPRC5A. In contrast, another antibody raised against the same antigen, HPA064241, was highly selective only for GPRC5D.

(GPRC5D Protein Recognition Site Ma)
Three of the six antibodies tested against corticotropin-releasing hormone receptor 1 (CRHR1) selectively recognized CRHR1 and associated with an antigen representing ICL1.

(CRHR1 Protein Recognition Site Map)
Next, the researchers analyzed the epitope properties that caused the antibodies to exhibit specific binding or not , based on all antigen structures predicted from AlphaFoldDB : the target antibodies were found to be significantly enriched in protein regions predicted to be disordered, as defined by low predicted local distance difference test (pLDDT) scores; in addition, the on-target epitopes in residues exposed to the surrounding environment were also enhanced, as defined by the average relative solvent accessible surface area (SASA).

(Antigen Distance Difference (pLDDT) and Relative Solvent Accessible Surface Area (SASA) Tests)
Conclusion:
After mapping the recognition sites based on polyclonal antibodies, specific monoclonal antibodies can be purified from the polyclonal mixture using the acquired sites. Following testing for specificity in expected determinations, the most suitable sites can be used to generate monoclonal antibodies. The paper also provides additional information on all antibodies and recognized sites, offering a design and screening strategy for guiding the development of GPCR target antibodies.
This paper intuitively demonstrates the limitations of anti-GPCR antibodies: they need to ensure stable specificity in recognizing GPCRs in different shapes and conformations, and they also need strong affinity properties. In addition, the exposed epitopes of GPCR proteins are scarce, making antibody development difficult. Rabbit antibodies have smaller antigen recognition epitopes and a rabbit-specific high-affinity maturation system, making them a good animal source for immunity.
