During antibody development, selecting the right antibody that binds to the correct epitope of an antigen is crucial for achieving optimal therapeutic efficacy and leveraging the desired mechanism of action:
For neutralizing or blocking properties, antibodies are often chosen to bind epitopes located at or near the interface between receptor and ligand;
For agonist antibodies, epitopes homologous to previously well-validated agonistic anti-mouse antibodies can be selected;
Regarding antibody effector functions, different antigen recognition epitopes can differentially recruit complement to exert complement-dependent cytotoxicity (CDC) function, affecting the level of antibody-dependent cell-mediated cytotoxicity (ADCC) induction;
In addition, antibodies binding to different antigenic epitopes may differentially affect receptor heterodimerization, internalization, ligand binding, and ultimately receptor function.
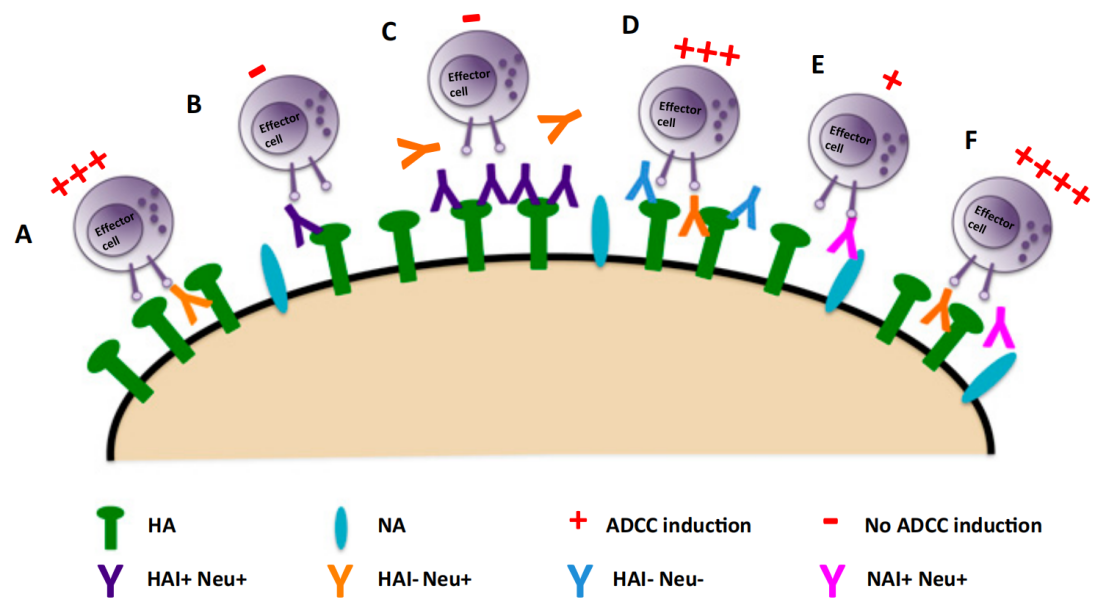
(Data source: He W, et al. Proc Natl Acad Sci USA. 2016)
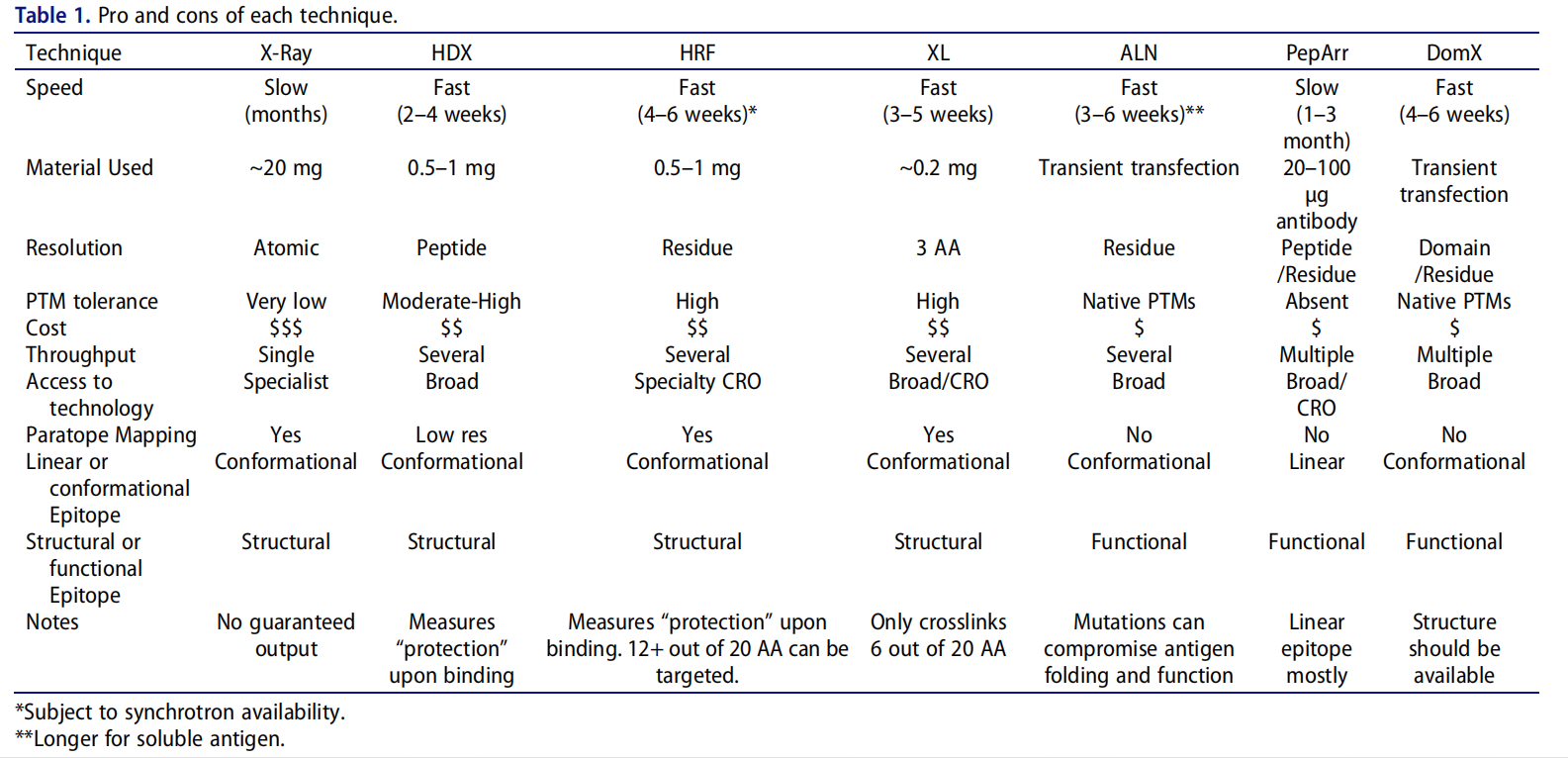
Recently, mAbs published a research paper titled“Epitope mapping of monoclonal antibodies: a comprehensive comparison of different technologies”, which systematically summarized seven widely used antibody epitope identification methods (X-ray crystallography, peptide arrays, alanine scanning, domain exchange, hydrogen-deuterium exchange, chemical cross-linking and hydroxyl radical imprinting), and explained the advantages and disadvantages of each technology.

(Article publication information)
X-Ray:
This method uses X-ray patterns diffracted by protein crystals to resolve the protein's atomic structure, revealing detailed information about the antibody-antigen interaction. X-rays are considered the gold standard for accurately identifying key interactions between antibodies and antigens at atomic resolution, with high precision and accuracy. Current state-of-the-art applications can resolve antibody-bound antigens to as high as 1.2 Å. However, obtaining the crystal structure of the antibody-antigen complex is time-consuming, has a low success rate, and is costly.

(Data source: Bianchi M, et al. Immunity. 2018)
Peptide array (PepArr):
This method utilizes a peptide library to screen peptides of varying lengths and sequences to identify specific peptide fragments that bind to antibodies, thereby revealing key regions of interaction between antibodies and antigens. The advantage of peptide microarray antibody epitope screening is that it allows for high-throughput analysis of interactions between antibodies and protein antigens in a relatively short period of time. However, peptides may not fully mimic the structure and function of the entire protein when used in antibody binding.
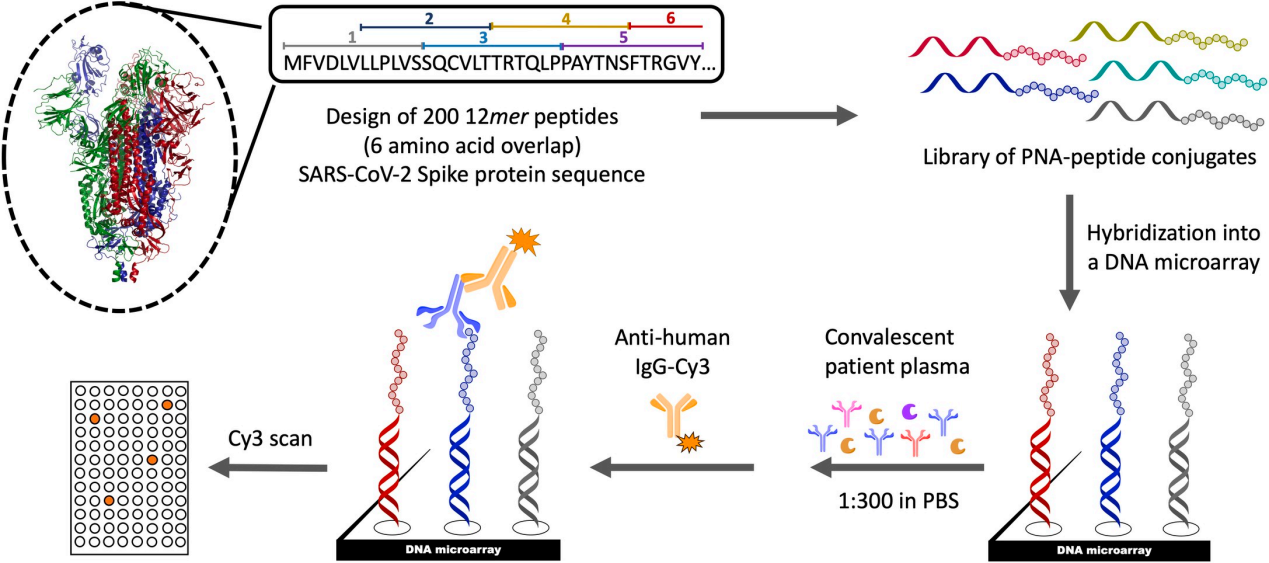
(Data source: Farrera-Soler L, et al. PLoS One. 2020)
Alanine Scan (ALN):
Alanine scanning utilizes the principle of alanine scanning, replacing amino acids in the antibody binding region one by one and evaluating the effect of each substitution on antibody-antigen binding. The experimental results are analyzed to identify amino acid residues that are critical for antibody binding, i.e., the composition of the antibody epitope. Alanine scanning can provide high-resolution information about the antibody binding site, helping to understand the details of the interaction between the antibody and the protein antigen. This method generally relies on precise structural information of the antibody and protein antigen, but alanine scanning does not take into account the overall structure and dynamics of the protein antigen.
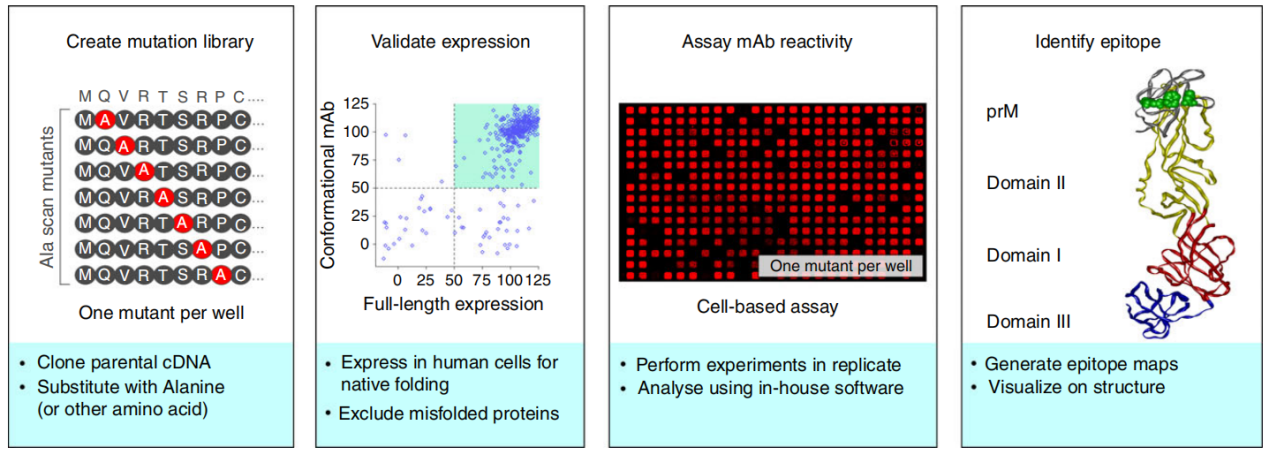
(Data source: Davidson E, et al. Immunology. 2014)
Domain exchange (DomX):
By replacing domains or fragments in an antigen with equivalent structural elements from homologous sequences in different species, the binding properties of these domain-swapped antigens and antibodies can be studied to infer key regions of antibody-antigen interaction. Domain swapping can only identify residues that naturally differ between species, and these residues may not play a role in the observed loss of binding.
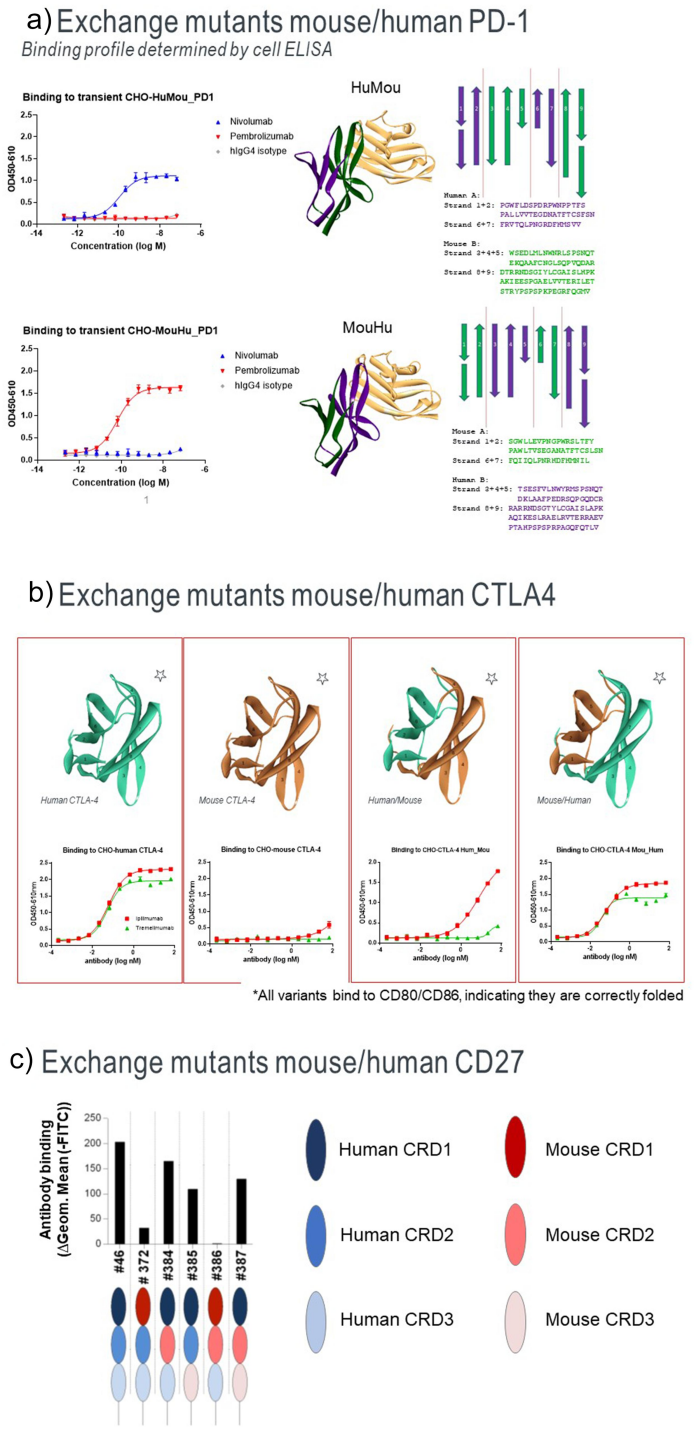
(Data source: Dang X, et al. MAbs. 2023)
Hydrogen-deuterium exchange (HDX):
Hydrogen-deuterium exchange (H2O) is a mass spectrometry-based epitope mapping technique. The extent of H2O exchange is determined by exposing the protein to deuterated water (D2O) in solution and then analyzing it by mass spectrometry. When an antibody binds to an antigen, it undergoes H2-deuterium exchange in the binding region, revealing the binding site. H2-deuterium exchange antibody epitope screening offers the advantages of high sensitivity and high resolution, allowing for understanding antibody-antigen interactions at the protein structural level. However, specialized mass spectrometry analysis is required, and interpretation of the results often requires in-depth expertise.
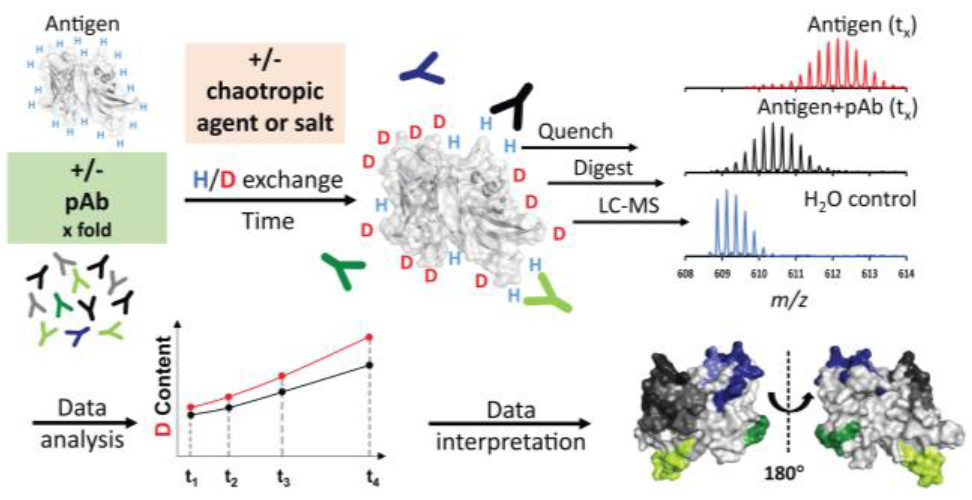
(Data source: Ständer S, et al. Anal Chem. 2021)
Chemical cross-linking (XL):
This method typically requires the use of specific chemical cross-linking reagents that form reversible or irreversible covalent bonds to bind antibodies to antigens. This method involves cross-linking the antibody to the antigen using a cross-linking reagent, and then analyzing the cross-linked product by mass spectrometry to identify the antibody binding site (epitope). Chemical cross-linking can provide direct information about the antibody binding site and is independent of the antibody's crystal structure. However, chemical cross-linking may result in some structural changes, and the selection of appropriate cross-linking reagents and conditions is crucial for the successful execution of this experiment.
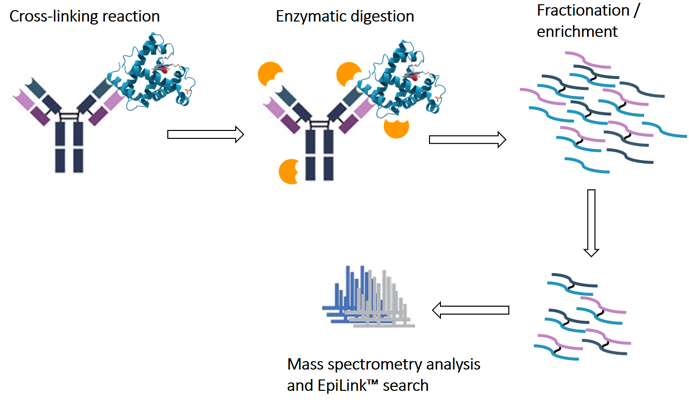
(Data source: MtoZ-Biolabs)
Hydroxyl Radical Signature (HRF):
Similar to the chemical cross-linking method, this method is based on the oxidative sensitivity of hydroxyl radicals to protein residues: the antibody and protein antigen interact to form an antibody-antigen complex, and hydroxyl radicals are introduced into the solution to react with the antibody-antigen complex to label the amino acid residues in the antibody binding region. The hydroxyl-labeled protein is then subjected to mass spectrometry analysis to determine the location of the hydroxyl label.
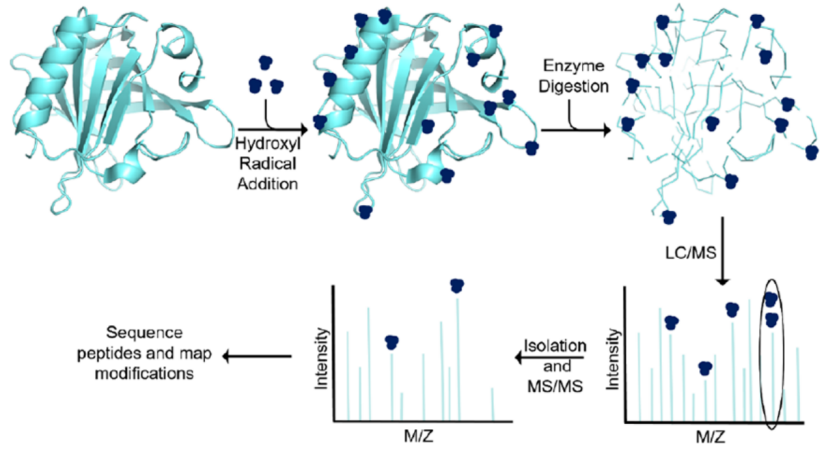
(Data source: McKenzie-Coe A, et al. Chem Rev. 2022)
Summary:

Three functional-based methods—domain exchange, peptide arrays, and alanine scanning—provide chain-, peptide-, and residue-level resolution, respectively. Peptide arrays have the lowest success rate of all the techniques tested in this publication. The value of domain exchange lies in its high success rate in producing high-quality mapping results, which can be cross-validated with other methods and used to guide residue selection in alanine scanning. Hydrogen-deuterium exchange, chemical cross-linking, and hydroxyl radical imprinting are all structure-based epitope mapping methods. All three techniques demonstrate a high probability of success and are not limited by antigen size. All require expensive high-resolution mass spectrometry and specialized interpretation expertise.