Background
Microscale thermophoresis (MST) is a biophysical technique that measures the strength of an interaction between two molecules by detecting changes in fluorescence signal caused by infrared laser-induced temperature changes. The range of the fluorescence signal change correlates with the binding of the ligand to the fluorescent target. In general, MST is based on the movement of fluorescent molecules within a temperature gradient. MST is widely applicable in the pM to M concentration range and requires minimal sample volume, only a few microliters. Furthermore, no target protein immobilization is required. You can either fluorescently tag the target protein or exploit the intrinsic fluorescence of the target. In either case, one binding partner is fluorescently labeled, while the other remains unlabeled. The fluorescently labeled molecule serves as a reporter molecule, making it easy to detect the change in the MST signal caused by binding. The results are automatically analyzed by software to accurately calculate the Kd value.
Principle: The core principle of MST is based on the phenomenon of thermophoresis. An infrared laser is used to generate a local temperature gradient in the solution, and the changes in the thermophoretic motion trajectory caused by the changes in the size, charge, hydration layer and other properties of the molecules due to binding are monitored through fluorescent labeling, thereby quantifying the interaction between molecules. Its main principles can be divided into the following parts: a) Temperature gradient drives swimming ; b) Molecular properties determine swimming behavior ; c) Binding causes property changes ; d) Detection of binding events : The MST trajectory changes significantly after the two molecules bind, and one of the fluorescently labeled molecules can be detected by high-sensitivity fluorescence to perform real-time monitoring ; e) Quantitative analysis : By measuring the change in the MST trajectory of the target molecule under different ligand concentrations, the binding curve can be drawn, and the precise binding constant (KD) can be calculated.
-2.png)
( Data from Plach MG, et al. Bio Protoc. 2017)
Experimental steps/process
(1)Fluorescent labeling : One of the two interacting molecules (A and B) is fluorescently labeled (usually the target molecule, such as a protein). The labeling agent must not affect the binding activity of the molecules. Common labeling methods include NHS chemical labeling and His-tag labeling with fluorescent dyes.
(2)Sample preparation : Prepare fluorescent labeled protein, set up protein stock solution (such as 50 μ L ), diluted 1:100 (add 4950 μ L buffer), take 10 Add μL of this dilution to each ligand dilution and incubate at room temperature (the time depends on the affinity and can be extended if the affinity is low ) .
(3)Sample loading and measurement : Load the incubated sample onto the instrument, start MST analysis, and detect the thermodynamic and kinetic parameters of intermolecular interactions.
(4)Data analysis: The electrophoresis parameters at different ligand concentrations were plotted against the ligand concentration to obtain the binding curve . The binding constant (KD), Hill coefficient, stoichiometric ratio and other parameters were calculated through nonlinear fitting.
-3.png)
(Data source Romain M , et al. Bio Protoc. 2020)
Features of MST:
(1) Ultra-high sensitivity: requires only trace amounts of sample (nanoliter volume, picomolar concentration), suitable for rare or difficult-to-purify biological molecules (such as membrane proteins); can detect weak binding (KD range covers pM~mM), and is particularly good at fragment-based screening.
(2) In situ detection in solution: a) No need for immobilization: molecules diffuse freely, avoiding false positives or conformational restrictions caused by solid phase carriers; b) Near-physiological environment: direct measurement in buffer, cell lysate, or even complex systems containing serum/detergents, the results are more physiologically relevant.
(3) Fast and efficient: A single measurement takes only a few minutes and can be analyzed in parallel with high throughput (such as in 96-well plate format), making it suitable for rapid screening.
(4) Universality: It is compatible with all types of biological molecules (proteins, nucleic acids, small molecules, ions, liposomes, etc.) and can study various interactions such as protein-protein, protein-nucleic acid, and protein-small molecules.
(5) Direct quantitative parameters: direct output of binding constant (KD), stoichiometric ratio, Hill coefficient, and some modes can also obtain kinetic parameters (kon/koff).
(6) Low sample consumption: Each measurement consumes only a microliter of sample, significantly reducing experimental costs.
Applications of MST:
1. Research on biomolecular interactions: protein-protein interactions (such as antigen and antibody); protein-nucleic acid interactions (transcription factors and DNA/RNA); protein-small molecule interactions (such as target protein and drug screening); nucleic acid-nucleic acid interactions (such as nucleic acid hybridization);
2. Drug discovery and development : such as high-throughput screening (HTS) validation, structure-activity relationship studies
3. Antibody characterization : Determine the binding affinity (KD) of the antibody to the antigen
4. Aptamer screening and characterization
-4.png)
Case:
Protein-Protein Interactions:
Zhou Rui et al., using GST pull-down assays, identified an interaction between the middle tail domain (Myo5a-MTD) of myosin-5a and the actin-binding domain (Mlph-ABD) of the cargo protein melanavidin (Mlph). Using isothermal titration calorimetry (ITC) or microthermophoresis (MST), they determined the dissociation constant between Myo5a-MTD and Mlph-ABD to be 562±169 nM. Ingvild Gudim et al. also measured the binding affinities of three different flavodoxin reductases (FNRs) and a flavodoxin-like protein (NrdI, from Bacillus subtilis) using microthermophoresis (MST). They demonstrated that NrdI is essential for activating the manganese-binding form of the class Ib ribonucleotide reductase (RNR) system. RNRs are the sole source of de novo deoxyribonucleotides required for DNA replication and repair in all organisms.
-5.png)
(Data source: Left: Rui Zhou , et al. Bio Protoc. 20 25; Right: Gudim I, et al. Bio Protoc. 2017)
Protein-nucleic acid interactions:
Gregory T et al. describe a protocol for obtaining binding curves of DNA aptamers to SARS-CoV-2 using enzyme-linked oligonucleotide assay (ELONA) and microscale thermophoresis (MST). These methods can be used to determine the binding affinity of aptamers to infectious SARS-CoV-2 and their selectivity against SARS-CoV-2 and other viruses that have been rendered infective by UV inactivation.
-6.png)
(Data source: Pawel GT , et al. Bio Protoc. 2022)
Protein- peptide interactions:
Plach et al. describe in detail two MST assays for analyzing ( 1 ) the interaction between a specific peptide from eukaryotic RNA polymerase II and a protein subunit of the eukaryotic transcription elongation complex, and ( 2 ) the interaction between an N-terminal histone tail peptide and an epigenetic “reader” protein. These experiments demonstrate that MST can characterize protein - peptide interactions that are triggered by small changes in the peptide (e.g., a single phosphorylation modification on a specific serine residue) .
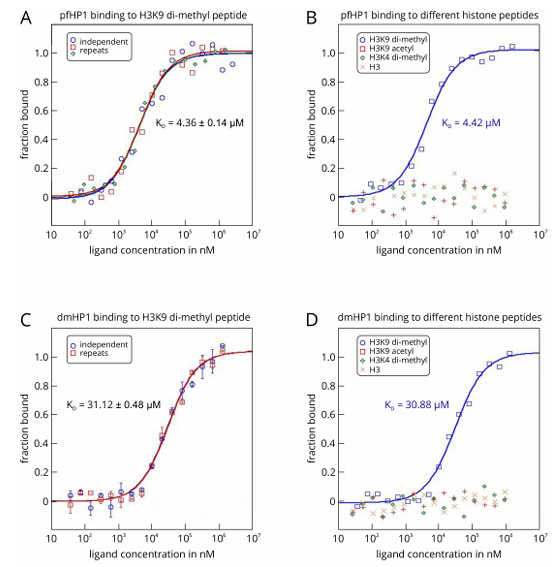
(Data from Plach MG, et al. Bio Protoc. 2017)
Protein- Small Molecule Interactions:
In 2019, Wu Dalei and others from Shandong University combined the research methods of crystallography, biophysics and MST to explore the chemical ligands of hypoxia-inducible factor- 2 / HIF-2α (a heterodimeric transcription factor formed by the dimerization of the oxygen-sensitive HIF-2α subunit and its essential partner subunit ARNT) and discovered its chemically unrelated antagonist (T1001). They use the same mechanism of action: after binding, they will inhibit HIF-2α. Residue M252 within the PAS-B pocket pushes toward the ARNT subunit, thereby weakening heterodimerization.
-8.png)
(Data from Wu D , et al. Nat Chem Biol . 2019)
Antigen-antibody interaction:
Microscale thermophoresis (MST) analysis of CR3022, an antibody against the S protein RBD developed by antibodies online, revealed an affinity (dissociation constant, KD) of 1.09 nM for the canonical SARS-CoV-2 S protein. MST was further used to test the affinity of CR3022 for the S proteins of different strains (B.1.1.7/Alpha, B.1.351/Beta, B.1.617.2/Delta, B.1.617.1/Kappa, and P.1/Gamma). The results showed that KD values for all tested variants were in the nM range, indicating that CR3022 maintains high affinity for all of them. (Note: Lower KD values indicate stronger binding.)
-9.png)
(Data source: Antibodies - online official website)
