Service Support
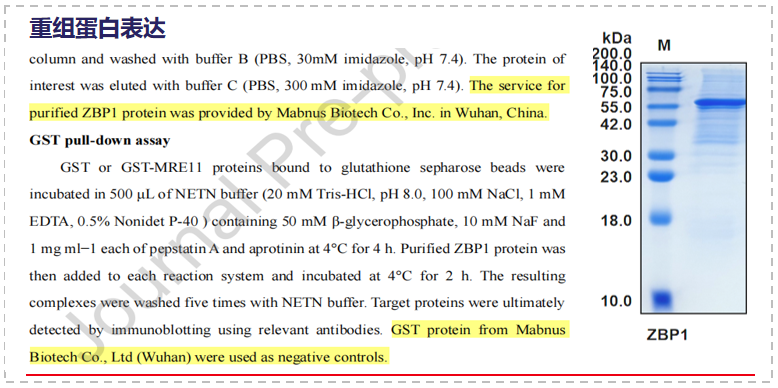
Mabnus Bio undertook the custom development of the active protein ZBP1 in this study.

Background

Genomic instability, a hallmark of cancer, stems from defects in the DNA damage response (DDR). In cancer cells, DNA double-strand breaks (DSBs) are the primary source of damage threatening genomic integrity, and DSB repair primarily occurs through two mechanisms: non-homologous end joining (NHEJ) and homologous recombination (HR). Various pattern recognition receptors (PRRs), including cGAS, AIM2, DHX9, RIG-I, and ZBP1, can recognize cytoplasmic nucleic acids (cNAs), activating cytoplasmic immune pathways, thereby inducing interferon (IFN) production and triggering pro-inflammatory responses. ZBP1 is a classic pattern recognition receptor (PRR) involved in regulating programmed cell death and innate immune responses. However, the role of ZBP1 in the cell nucleus remains unclear. It is currently unknown whether ZBP1 affects the DNA damage response (DDR) in a similar manner to other pattern recognition receptors.

On October 28, 2025, the Department of Obstetrics and Gynecology team at Tongji Hospital Tongji Medical College of HUST, published a paper entitled "ZBP1 antagonizes MRE11-mediated DNA end resection and confers synthetic" in Drug Resistance Updates. This study revealed the regulatory mechanism of ZBP1 in DNA double-strand break (DDR) repair. When exogenous DNA damage is induced, nuclear ZBP1 accumulates at the double-strand break site and is phosphorylated by ATM kinase at serine 106. Phosphorylated ZBP1 then binds to MRE11, preventing downstream end-excision reactions and thus blocking homologous recombination (HR) repair. The study found that high ZBP1 expression levels in ovarian cancer patients were significantly correlated with sensitivity to the synthetic lethal effect of poly(ADP-ribose) polymerase (PARP) inhibitors, and these drugs were more effective in HR-deficient tumors.
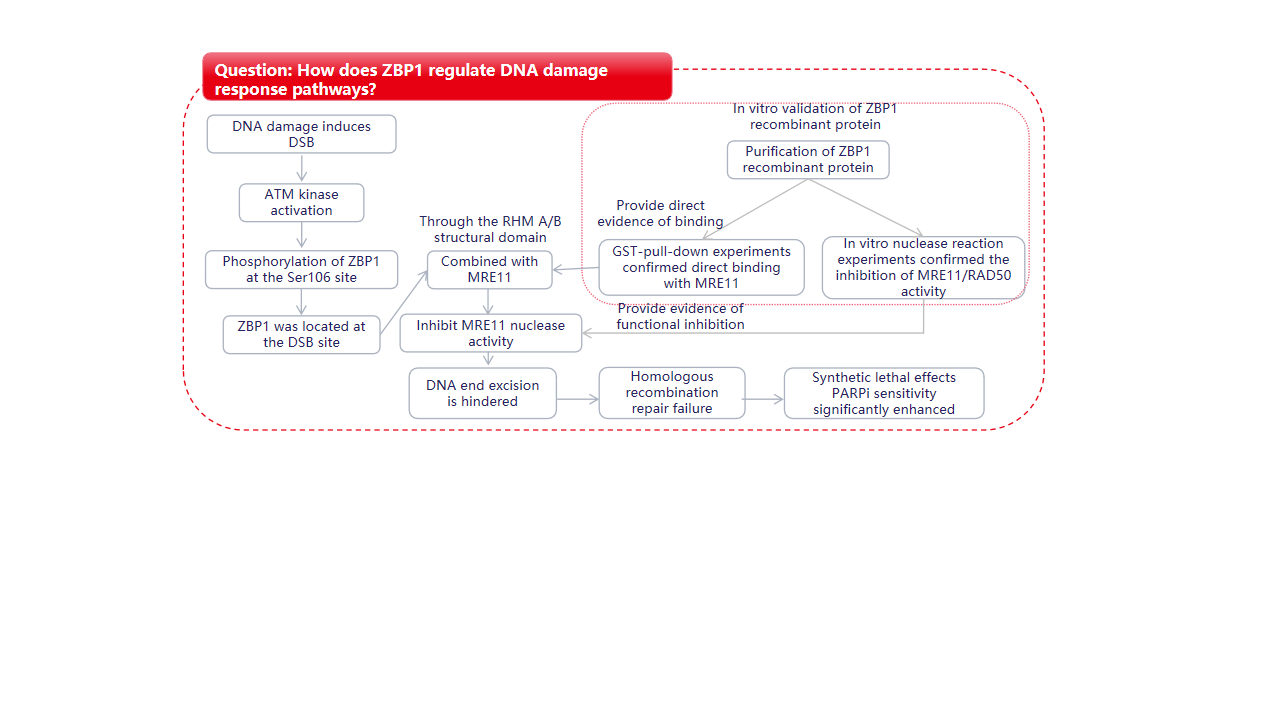
Research Results
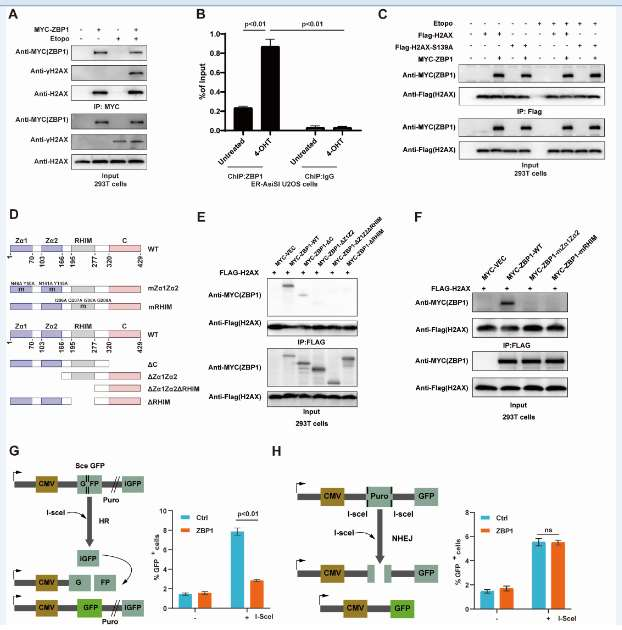
Experiments including co-immunoprecipitation (Co-IP), chromatin immunoprecipitation (ChIP), construction of ZBP1 truncated variants and point mutations, and immunofluorescence showed that ZBP1 participates in and negatively regulates DNA double-strand breaks (DSB). ZBP1 is recruited to DSB sites and specifically inhibits the HR repair pathway. ZBP1 blocks HR by inhibiting DNA end excision.
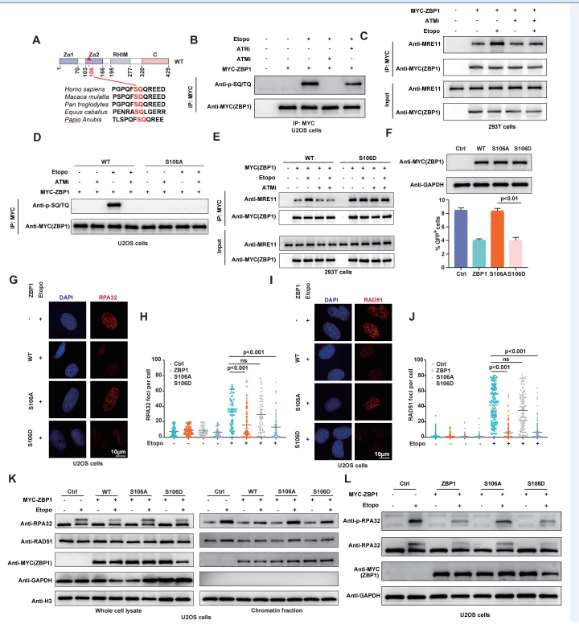
Through Co-IP, GST pull-down, and in vitro nuclease activity assays, it was found that ZBP1 inhibits DNA end excision by directly binding to MRE11 and inhibiting its nuclease activity. ATM-mediated phosphorylation of ZBP1 regulates its binding to MRE11 and its HR inhibitory function.
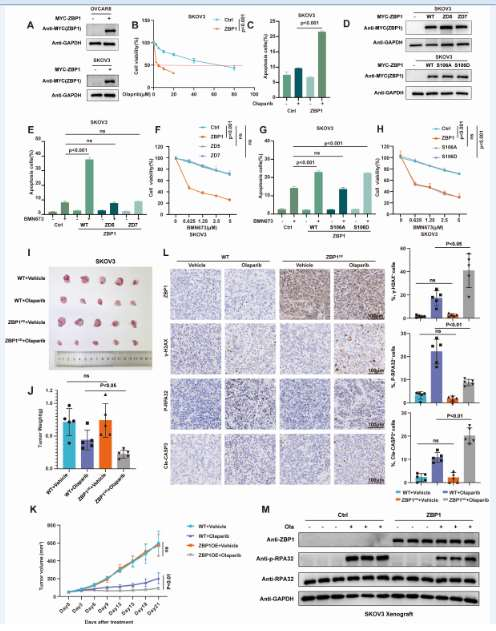
By overexpressing ZBP1 in ovarian and breast cancer cells, CCK-8 cell viability and Annexin V flow cytometry apoptosis were detected; a mouse subcutaneous xenograft model was established. It was found that ZBP1, by inhibiting heart rate (HR), conferred synthetic lethality to tumors against PARP inhibitors both in vitro and in vivo. ZBP1 may serve as a potential biomarker for predicting the response of ovarian cancer patients to PARP inhibitor therapy.
Summarize
Following DNA damage, ZBP1 is phosphorylated by ATM, which then directly binds to MRE11 and inhibits its nuclease activity, ultimately suppressing DNA end excision and HR repair. This mechanism not only explains why high ZBP1 expression enhances the efficacy of PARP inhibitors, but more importantly, it establishes ZBP1 as a potential biomarker for predicting PARP inhibitor response, providing a new theoretical basis and clinical strategy for precision treatment of ovarian cancer.

