Background
Antibodies are essential components of the immune system and have a wide range of biomedical applications. Elucidating the complex interactions between antibodies and antigens is a crucial step in drug development. However, the complexity and extensive nature of the data present significant challenges in accurately identifying and understanding these interactions. To overcome these challenges and deepen our understanding of the antibody-antigen interface, Professor Huang Jian's research team at the University of Electronic Science and Technology of China developed the Antigen-Antibody Complex Database (AACDB).

On May 22, 2025, Professor Huang Jian's research team published an article titled "A comprehensive antigen-antibody complex database unlocking insights into the interaction interface" in Elife. The article provides a detailed introduction to the AACDB, a database that integrates data on antibody developability and antigen-drug target relationships, and is of great value in assisting the development of new antibody therapeutics. The database contains comprehensive information on antibody-antigen epitopes, making it a valuable benchmark in immunoinformatics research. AACDB also provides detailed information on interacting residues using two methods (ΔSASA and atomic distances), facilitating benchmarking of epitopes and epitope prediction methods. AACDB provides a user-friendly interface for querying, manipulating, browsing, and visualizing comprehensive information on antibody-antigen complexes. It also offers customized datasets to meet specific research needs. Researchers can access AACDB fully online (http://i.uestc.edu.cn/AACDB) and the database is regularly updated.
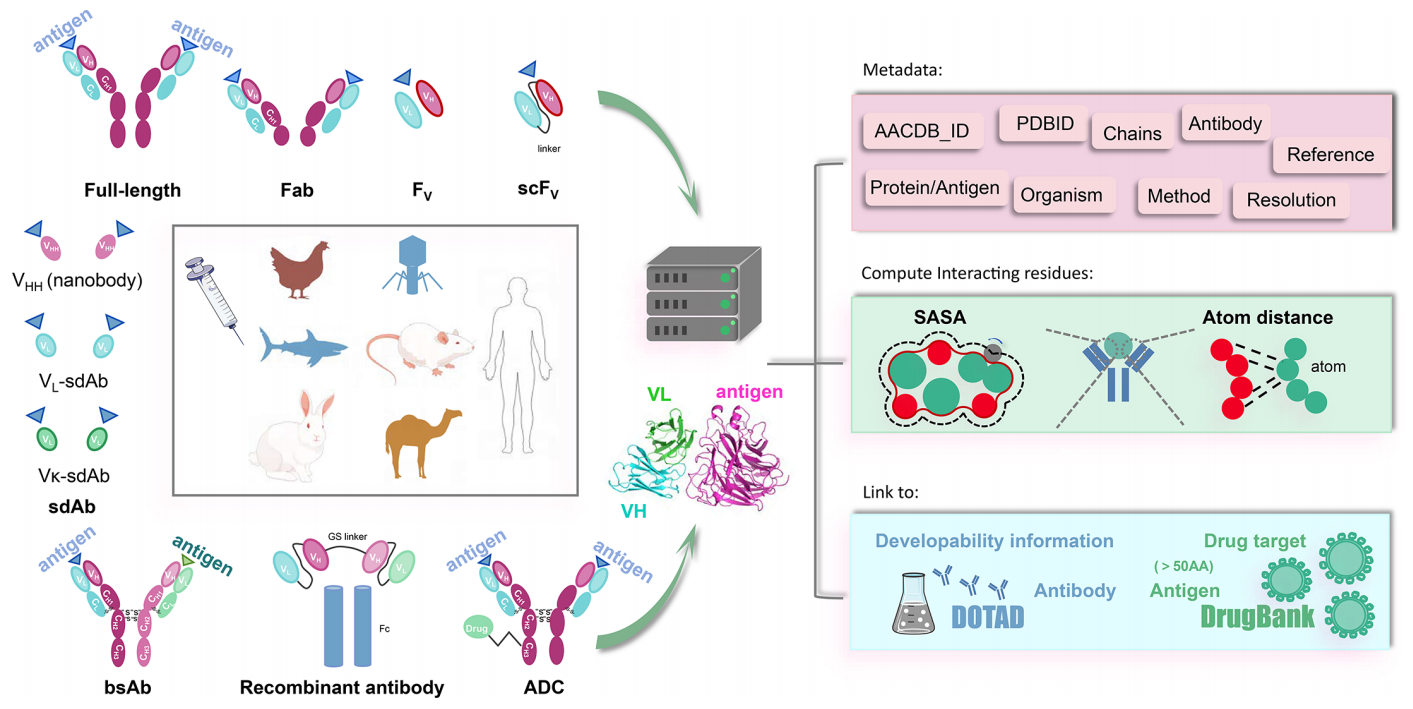
AACDB interface introduction
Data statistics interface
This interface mainly lists the distribution of antibody types in the database , the number of antigen-antibody complexes published in different years (unique PDBID) , and the distribution of antibody entries by organism.

Database browsing and search interface
This interface lists AACDB_ID, PDBID, light and heavy chain IDs , antibody, antigen, species origin, structural experimental method, protein structure resolution, and reference related information.
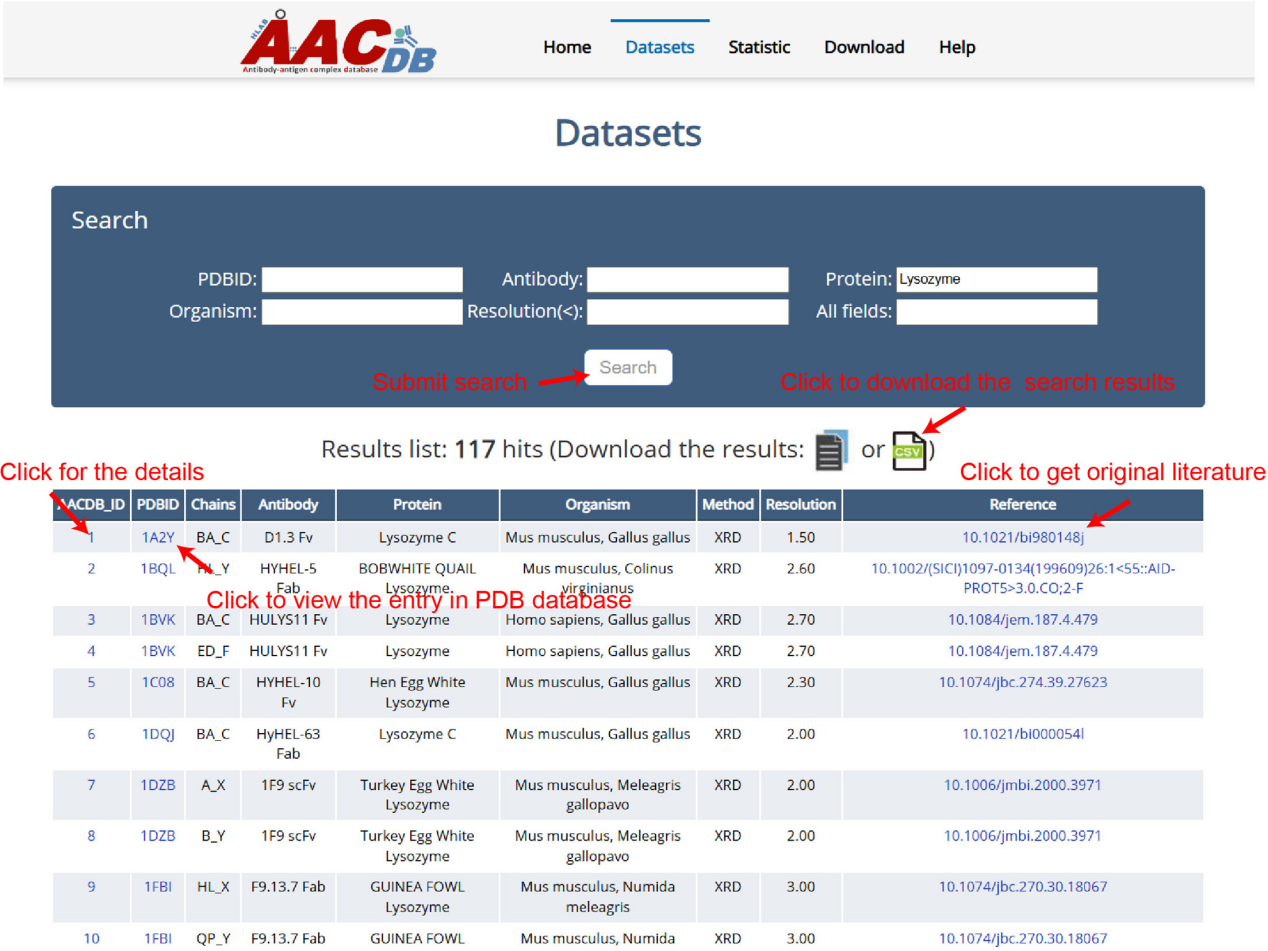
Individual structures can be accessed using the AACDB_ID. Clicking on the AACDB_ID will lead to a more detailed interface, including the antibody's mutations, INN and clinical trials, and the antigen's DrugBank ID (if available). Further details about each chain can be found under the Structure Information tab. These include: sequence, type and position of mutated amino acids, interacting residues in each chain based on the ΔSASA method, and an interaction map of binding site-epitope residues using a distance threshold of <6 Å. Data can be downloaded for each entry.

Advantages and limitations of AACDB
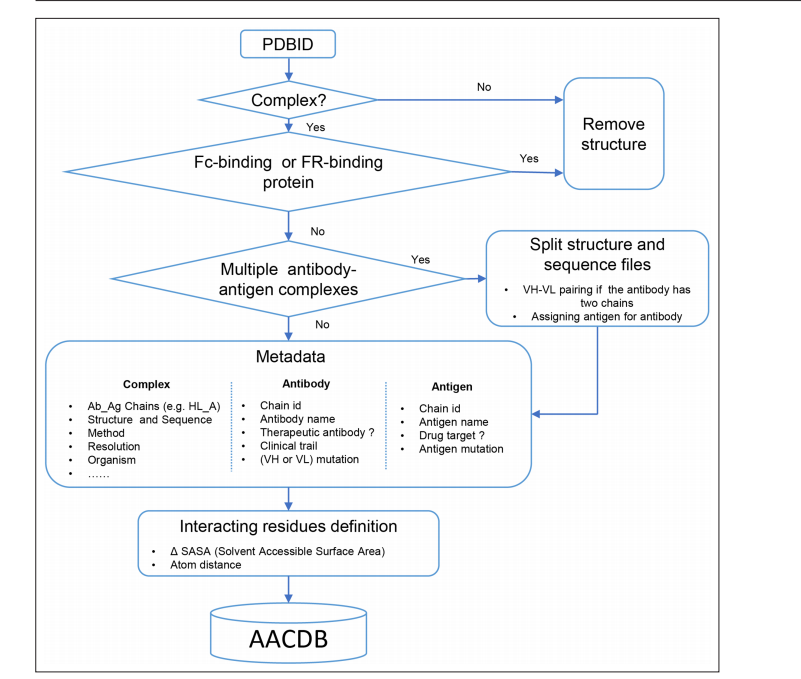
During the data curation, many errors in the PDB database annotations were discovered, and a lot of effort and time were invested in manually cross-referencing the original literature to correct these errors and rule out cases where SabDab incorrectly annotated antibody-binding proteins as antigens.
In addition to curating and re-annotating structural data, AACDB also provides features not found in other antigen-antibody complex databases: (1) AACDB's data processing pipeline supports mmCIF files; (2) amino acids on the interaction interface are provided through two methods, enabling the definition of unified standards for epitopes and paratopes. This provides a more accurate and comprehensive benchmark dataset for developed interaction interface prediction tools, enhancing the comparability of various tools.
Currently, the AACDB only includes antigenic proteins longer than 50 amino acids. The current version of the AACDB only includes antigenic proteins longer than 50 amino acids. Future iterations will expand structural coverage to include diverse antigenic classes (peptides, nucleic acids, haptens), while also incorporating literature-verified affinity data through systematic cross-referencing.
In conclusion
AACDB is a novel database of antigen-antibody complexes that provides information on antibody developability, antigen-drug-target relationships, and detailed antigen-antibody interaction interfaces. It is fully accessible at http://i.uestc.edu.cn/AACDB. We are committed to regularly updating the database to ensure that researchers in immunoinformatics have timely access to this valuable resource.
