Background
Deep learning has greatly advanced the design of highly stable static protein structures. However, mimicking the controllable conformational dynamics of natural switch proteins (with controlled conformational changes/dynamics required for signal transduction) remains a challenge that cannot be designed from scratch. This study proposes a general deep learning-guided method that achieves dynamic changes of proteins with atomic-level precision, which can be used to design dynamic changes between different conformations within protein domains from scratch.
In July 2024, Amy B. Guo of the University of California, San Francisco, published a research paper titled "Deep learning-guided design of dynamic proteins" in Science. Using a deep learning-guided approach, the paper resolved microsecond transitions between conformations of four structure-validated designs and demonstrated that the conformational landscape can be manipulated by both orthosteric ligands (e.g., enzyme catalytic pockets, receptor ligand binding sites, and other naturally functional core regions of proteins ) and allosteric mutations (e.g., surface residues and indirect contact regions, distant from the core functional regions). a) Orthosteric ligands can directly bind to functional regions, altering local conformation; b) Allosteric mutations can indirectly influence them through residue interaction networks (e.g., long-range coupling via hydrogen bonds and hydrophobic stacking). Physics-based simulations closely aligned with deep learning predictions and experimental data, revealing distinct state-dependent residue interaction networks and predicting mutations that tune the designed conformational landscape. Their approach demonstrates that novel motion patterns can now be achieved through de novo design, providing a framework for constructing bioinspired, tunable, and controllable protein signaling behaviors.
Biological functions of protein conformational changes
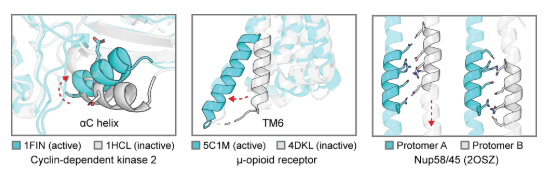
In natural proteins, conformational changes within domains are very common and are crucial for functional regulation, including molecular recognition, signal transduction, and enzyme catalysis . The number of conformational states and their transitions are determined by the relative free energy of the energy landscape and the potential barriers between them. Partners, the environment, post-translational modifications, and allosteric effectors can also regulate function by manipulating active/inactive conformational populations, and computationally designed mutations can also transform conformational landscapes and functions. For example, it is known that helices undergo various motion modes such as twisting, bending, and sliding to achieve different functions: such as forming active catalytic sites (such as the rotation of the αC helix in kinases, left figure below, Cα atomic root mean square deviation (RMSD) = 6.4Å), binding to downstream signaling molecules ( such as the bending of the TM6 helix in G protein-coupled receptors (GPCRs) to allow G protein coupling, in the figure below , Cα RMSD = 6.9Å), and regulating the pore diameter ( such as the sliding of the helix in nucleoporins, right figure below , Cα RMSD = 5.7 Å).
With tunable two-state equilibrium
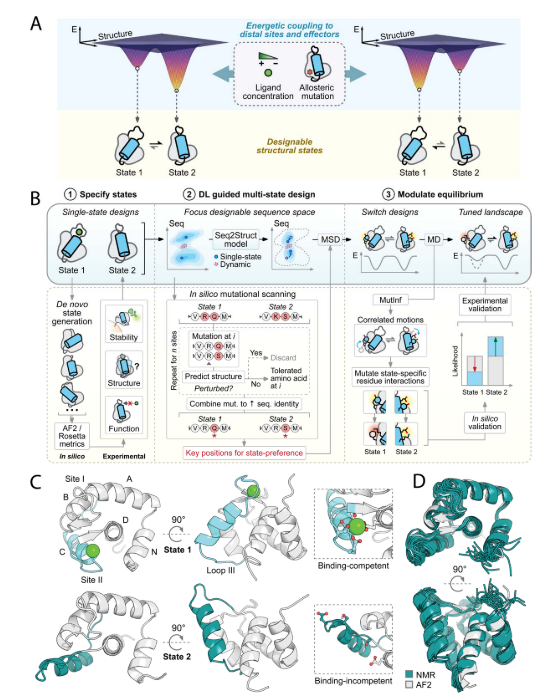
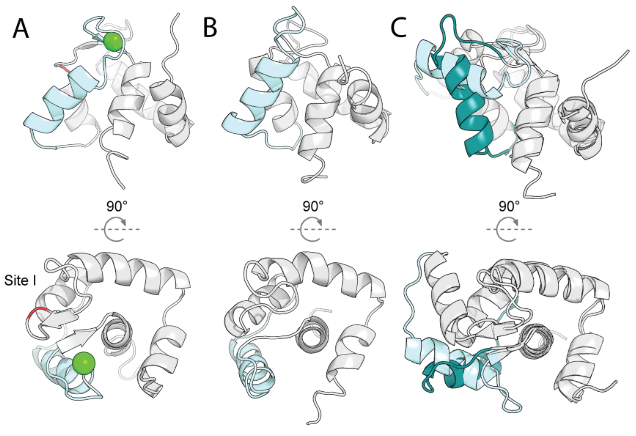
This study developed a general design strategy for constructing protein sequences with multiple energy minima, each corresponding to a specific conformational state (Figure A below). The design focused on intradomain conformational rearrangements of secondary structural elements (mimicking the core mechanism of natural signaling proteins) and achieved conformational equilibrium control through orthogonal ligands (binding to regions of conformational change) and allosteric perturbations (acting at distal sites). The design process consists of three stages (Figure B below): a) generating a library of candidate states through physical-based conformational sampling and evaluating the designability of each state; b) using deep learning to search sequence-structure space, identifying key residues and designing multi-state compatible sequences; c) integrating simulation with deep learning-based prediction of state-specific networks to screen for mutations that can modulate conformational switching. This study used an engineered Ca²⁺-binding protein (derived from the CN-terminal domain of troponin) as a proof-of-concept (Figure C below). The starting state was the E41A mutant (PDB: 1SMG), whose site I affinity was reduced to millimolar levels while site II maintained micromolar affinity. The mutant itself exhibited no Ca²⁺-dependent conformational changes. This approach demonstrates the general applicability of introducing controlled conformational changes into native proteins and also allows for the de novo design of binding-competent states.
De novo design of alternative states for generation and experimental verification
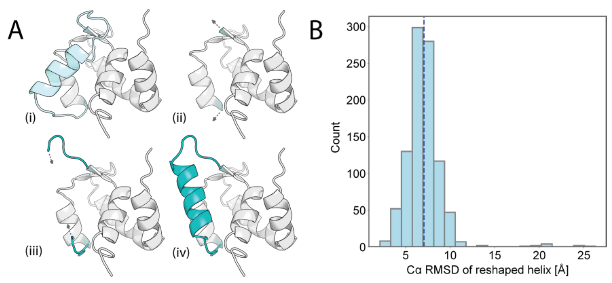
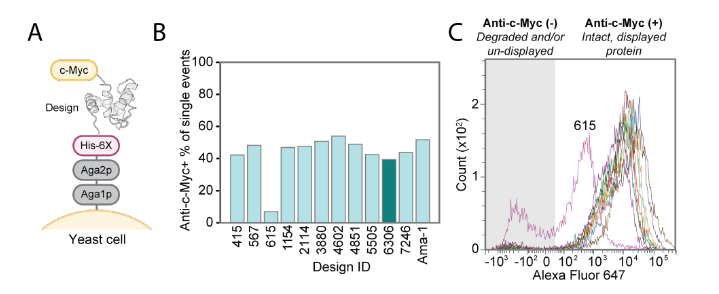
This study used the LUCS algorithm to sample a protein fragment containing loop III, helix C, and Ca²⁺ binding site II, generating a State 2 conformation that is geometrically distinct from State 1. This sampling allows for dynamic secondary structure transitions (loop-helix swaps), mimicking native conformational changes. Approximately 1,000 candidate models (average Cα RMSD 7.1 Å) were screened using Rosetta single-state sequence design. Eleven designs were experimentally validated, with 10 (such as #6306) being stable in yeast display. The NMR structure of design #6306 closely matched the AlphaFold2 prediction (Cα RMSD 0.98 Å). The displacement of helix C in design #6306 resulted in a reorganization of the Ca²⁺ binding site II and loss of binding ability, successfully achieving the design goal of an unbound state (State 2).
Polymorphic Design of Dynamic Proteins
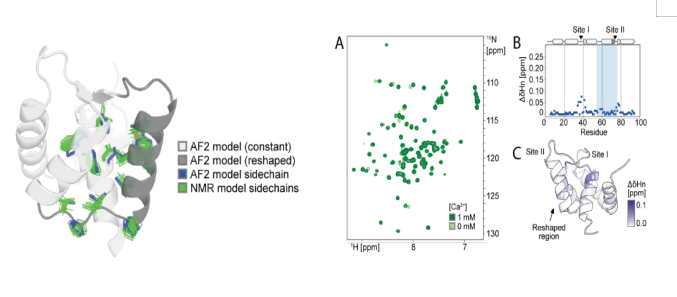
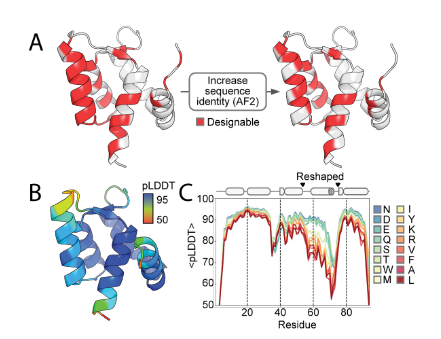
This study aimed to design polymorphic sequences that are compatible with both protein conformational states 1 and 2. AlphaFold 2 was used to screen key residues (such as solvent-accessible regions and hydrogen bond network sites), reducing the number of designable residues from 37 to 25. Combining protein MPNN generated highly similar sequences compatible with both states (only 15-18 mutations). Residue 89 was identified as the allosteric core: small hydrogen bond donors/acceptors (such as Ser) promote state 2 by stabilizing the loop III-helix D interaction; large hydrophobic residues (such as Leu) push away from loop III and prefer state 1. pLDDT analysis of AF2 confirmed that it can regulate distal conformations. This work achieved for the first time the precise control of protein polymorphic conformational distribution by fine-tuning the chemical properties of a single residue (89), providing a new approach for dynamic protein engineering.

Allosteric modulation of the conformational landscape by mutations
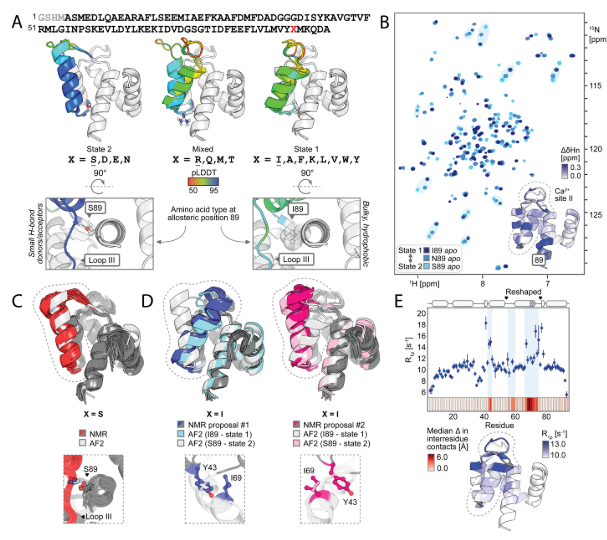
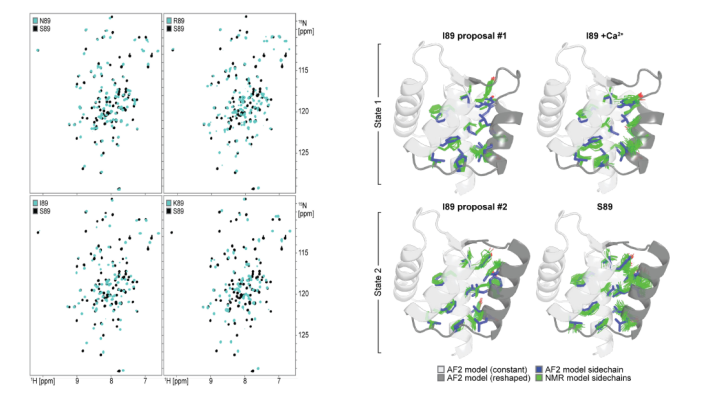
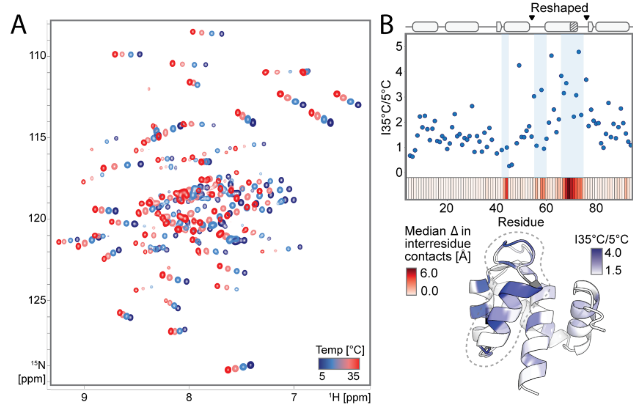
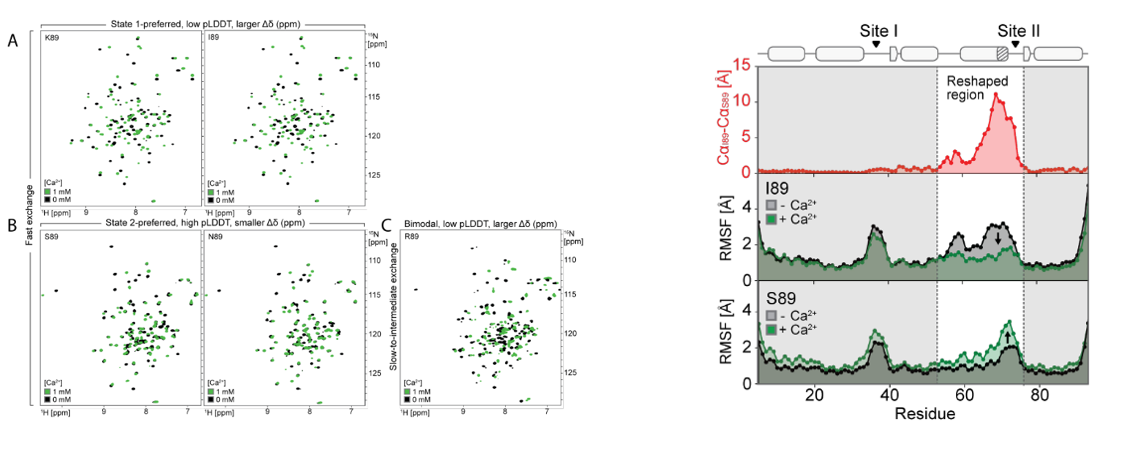
To verify whether the designed proteins form two predetermined conformations dependent on amino acid position 89, and to investigate the dynamics of these conformations, the researchers selected five designs (covering the AF2-predicted behavior of "preferred state 2 (S89, N89), preferred state 1 (I89, K89), and mixed state (R89)"). Using NMR spectroscopy, they measured ¹H-¹⁵N HSQC spectra and analyzed chemical shift differences. They also resolved the NMR structures of S89 (preferred state 2) and I89 (preferred state 1) and compared them with the AF2 predictions. They also collected temperature-dependent spectra of I89 from 5–35°C and analyzed the conformational exchange dynamics using rotating coordinate relaxation dispersion (R2). Finally, it was found that a single residue mutation at position 89 (such as I89→S89) caused a drastic change in the chemical shifts of 67/92 main-chain amides in the remodeled region. The displacement of N89 was between two states, and the remodeled region of I89 underwent two-state exchange on a low microsecond time scale. The temperature-dependent peak intensity changes and R2 values supported this dynamic equilibrium.
Orthosteric modulation of the conformational landscape by ligand binding
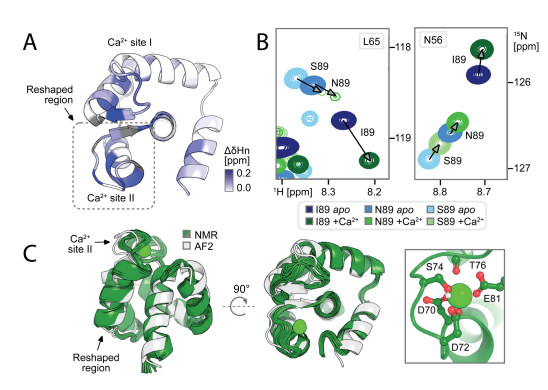
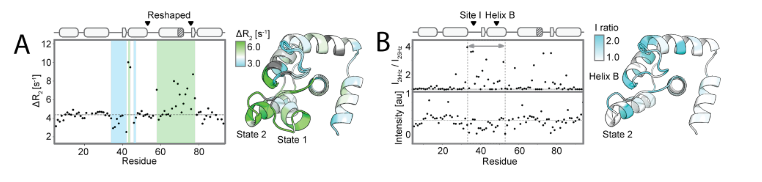
To explore whether Ca 2+binding also regulates the state population by preferentially stabilizing state 1, the researchers recorded 1H, 15N-HSQC spectra of each point mutant in the presence and absence of Ca 2+. By analyzing the NMR structure of I89 under Ca 2+ binding (holo structure), they ultimately found that (1) Ca 2+ induced significant chemical shift perturbations in the remodeling region and adjacent residues, and the direction was consistent with the equilibrium shift toward the bound state (state 1); (2) the holo I89 structure was highly consistent with the state 1 model, but residual dynamics of state 2 were still observed. The results indicate that the designed sequence can form two conformations in solution, and the abundance can be regulated by allosteric mutations and Ca 2+ binding.
Integrating physics-based simulations to tune conformational equilibrium
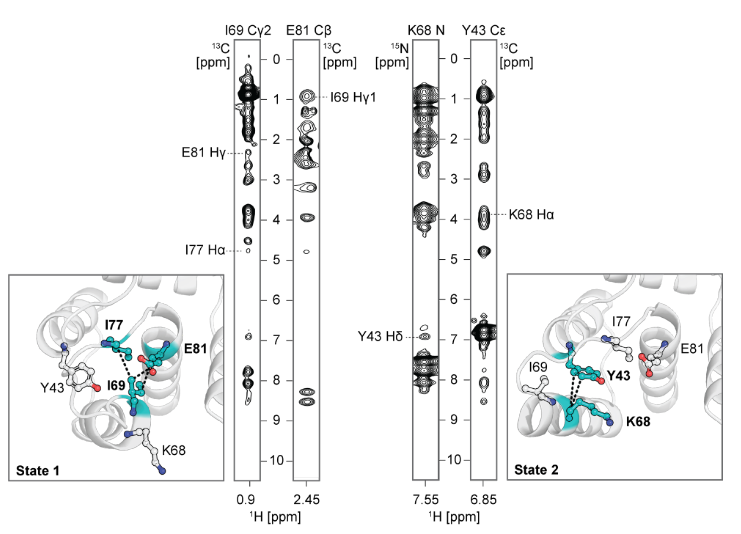
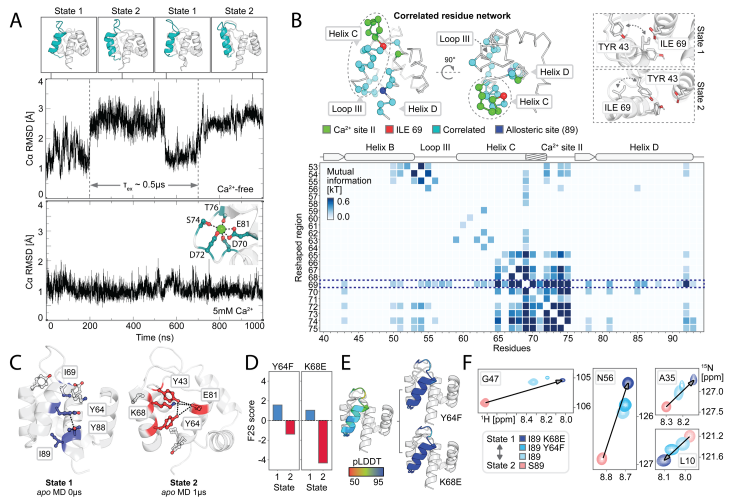
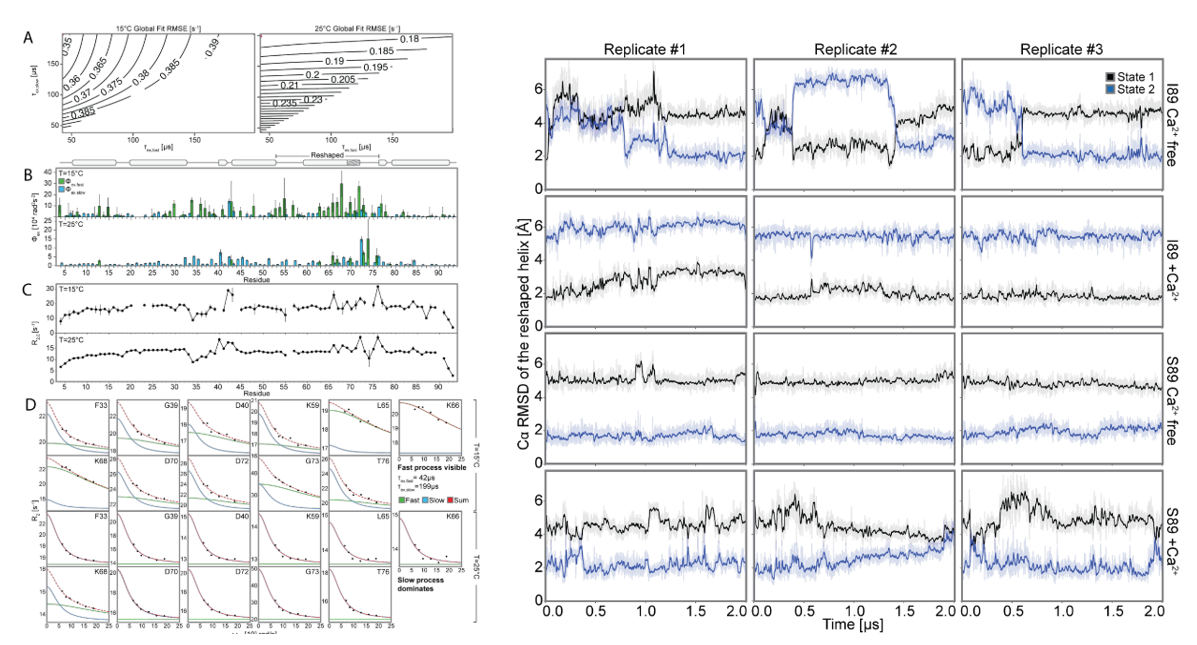
To further explore the atomic-level interactions behind the designed conformational transitions, the researchers performed molecular dynamics (MD) simulations on the designed I89 (with or without Ca 2+ ) and S89 (with or without Ca 2+ ) with and without Ca 2+ to analyze the conformational switching. The MD trajectory of apoI89 was analyzed to analyze the mutual information of the side chain torsional dynamics and the hydrophobic network. Mutations were predicted using the Frame2seq model, combined with AF2 evaluation, and then the conformational population changes were verified by NMR spectroscopy. The results showed that: (1) I89 switched between two states (state 1/2) at a low microsecond scale in the absence of Ca 2+ (consistent with R 2 ρ data); after adding Ca 2+, the Ca 2+ binding site II stabilized in state 1; (2) S89 remained in state 2 regardless of the presence of Ca 2+ (consistent with the high confidence of AF2 and experimental data). The mechanism is explained as follows: (1) a hydrophobic network connects Ca 2+ binding site II, loop III, and helix D, with position 89 differentially coupling this network through steric/hydrogen bonding; (2) a state-specific network is identified: in state 1, ILE69 is buried and TYR64/TYR88 are stacked in a π-π manner; in state 2, ILE69 is exposed and TYR43 is involved in hydrogen bonding/electrostatic interactions. The results explain the observed allosteric modulation of the conformational landscape by mutations at position 89 and validate predictions for further tuning the on-off balance.
Summarize
This study establishes a novel strategy for designing dual-conformation proteins: by synergistically regulating conformational interconversion through orthosteric ligands and allosteric mutations , it overcomes the limitations of traditional "domain replacement/rigid contact" design and realizes a novel model for kinase/GPCR-like interactions. Combining deep learning (design space compression and key sequence prediction) with physical simulations , the proposed method resolves a dual-state stable residue network. While unable to predict time-varying exchange, it can handle complex interactions between states. The successful generation of naturalistic conformational changes demonstrates the first "de novo design of allosteric switches" and the control of state abundance, demonstrating the ability to construct orthogonal switch and signaling systems, providing a universal tool for dynamic protein design and mechanism analysis.
