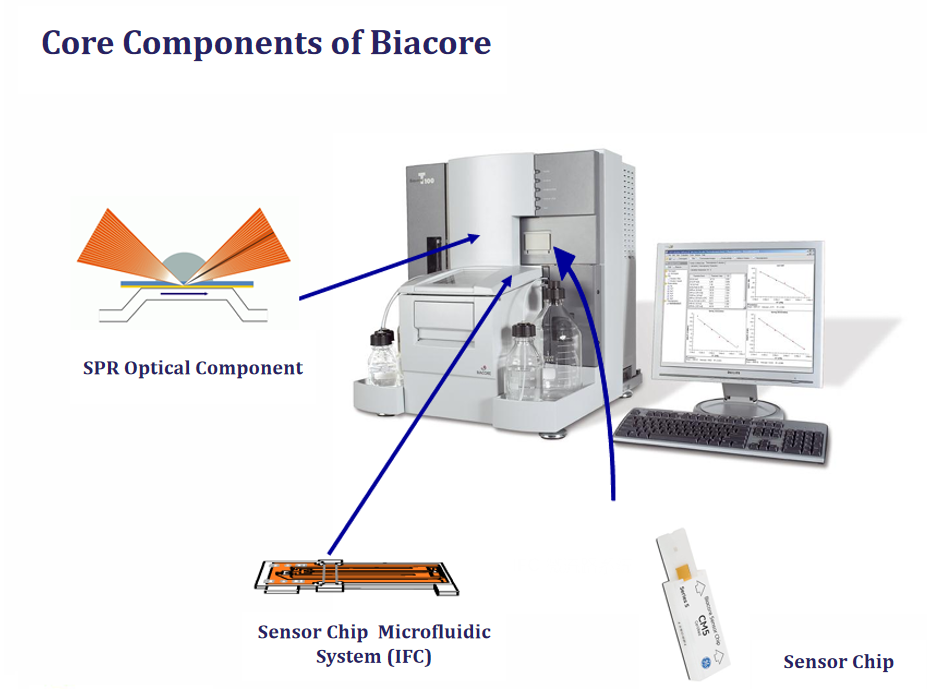

Background
Surface plasmon resonance (SPR) is a well-established, label-free technique for directly measuring biomolecular affinity. SPR analysis requires immobilizing an interacting component (ligand) on a conductive metal (primarily gold or silver) and continuously flowing a solution containing a potential binding partner (analyte) over the surface. The SPR phenomenon occurs when polarized light excites electrons at the interface of the metal and dielectric medium, generating electromagnetic waves that propagate parallel to the surface. The reflected light is detected to measure the change in refractive index caused by the interaction between the ligand and analyte, providing real-time data on kinetics and specificity.
Principle: The core principle of SPR technology is a physical optical phenomenon used for real-time, label-free detection of biomolecular interactions. Physical phenomenon: When a beam of polarized light strikes the interface between a metal film (usually a gold film) and a medium (such as a buffer) at a specific angle (incident angle), the light undergoes total internal reflection, generating an evanescent wave (Evanescent Wave) that penetrates the metal film. Plasmon resonance: The Evanescent Wave interacts with free electrons in the metal film, generating an electron density wave (called a surface plasmon wave, SPW) that propagates along the metal surface. Resonance condition: When the wave vector of the Evanescent Wave matches that of the Surface Plasmon Wave, energy transfer occurs, causing a sharp drop in the intensity of the reflected light, forming a resonance dip (SPR dip). The corresponding incident angle at this point is called the SPR angle. Detection: The SPR angle is extremely sensitive to the refractive index of the metal surface medium. Any binding event on the metal surface (such as protein-ligand binding) results in a change in the mass or concentration of surface material, resulting in a small shift in the refractive index. This shift immediately causes a shift in the SPR angle. By tracking the changes in the SPR angle (response signal, Response Unit, RU) in real time, the entire process of molecular binding and dissociation can be monitored in real time and dynamically without any labeling of the sample (such as fluorescence or radioactive labeling).
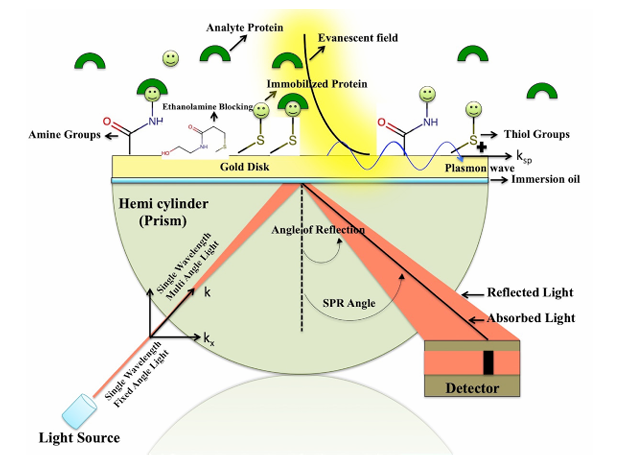
(Data source: Rath PP, et al. Bio Protoc. 2020)
Experimental steps/process
1) Sensor chip pretreatment: Select a suitable chip (such as carboxydextran chip CM5) and use reagents such as EDC/NHS to activate the carboxyl groups on the chip surface to convert them into active esters.
2) Ligand Immobilization: Known biomolecules (proteins, DNA, etc., referred to as "ligands") are immobilized on the chip surface through covalent coupling (e.g., amine coupling, thiol coupling, etc.). One flow channel immobilizes the ligand, while the other flow channel serves as a reference to subtract nonspecific signals.
3) Baseline balance: Load the incubated sample onto the instrument, start SPR analysis, and detect the thermodynamic and kinetic parameters of the intermolecular interaction.
4) Binding Assay: The sample molecules to be tested (called "analytes") are passed over the chip surface at a constant flow rate. a) Association Phase: The analyte binds to the ligand, increasing the mass of the chip surface and the SPR signal (response units, RU). b) Dissociation Phase: The sample is switched back to running buffer, dissociating the analyte from the ligand and causing the SPR signal to decrease.
5) Regeneration: Use mild acid, base or high salt solution to elute the still tightly bound analytes, so that the chip surface returns to its original state for the next injection.
6) Data analysis: The SPR signal changes throughout the entire process were monitored in real time (sensor graph), and the binding kinetic constants (association rate constant ka, dissociation rate constant kd) and affinity constant (KD) were obtained through software analysis.
SPR features:
1) Real-time monitoring/label-free: The dynamic process of molecular interactions can be observed in real time without fluorescent or radioactive labeling of samples.
2) High sensitivity: Able to detect very small mass changes (picogram level).
3) Provides rich kinetic information: not only the strength of binding (affinity, KD), but also the speed of binding (kinetics, ka, kd).
4) Low sample consumption: The required sample volume is very small (usually μL level).
5) Automation and high throughput: Modern SPR instruments enable automated operation and high-throughput screening.
6) Expensive instruments: The cost of equipment and consumables (such as chips) is high.
7) High requirements for buffer matching: The composition of the analyte solution and the running buffer must be highly consistent, otherwise huge false signals will be generated due to the difference in refractive index.
Applications of SPR:
1) Biomolecular interaction research: identification and quantitative analysis of protein-protein, protein-peptide, protein-nucleic acid, protein-small molecule, antibody-antigen and other interactions.
2) Drug screening and development: a) Lead compound screening: Rapidly screen small molecule drugs that bind to target proteins; b) Antibody drug characterization: Accurately measure the affinity and kinetics of antibodies for target antigens.
3) Epitope Mapping: Determine whether different antibodies bind to the same antigenic site.
4) Clinical diagnosis and biosensing: As a highly sensitive biosensor, it is used to detect disease markers, viruses, pathogens, etc.
5) Food safety and environmental monitoring: detecting allergens, toxins, pesticide residues, etc. in food.
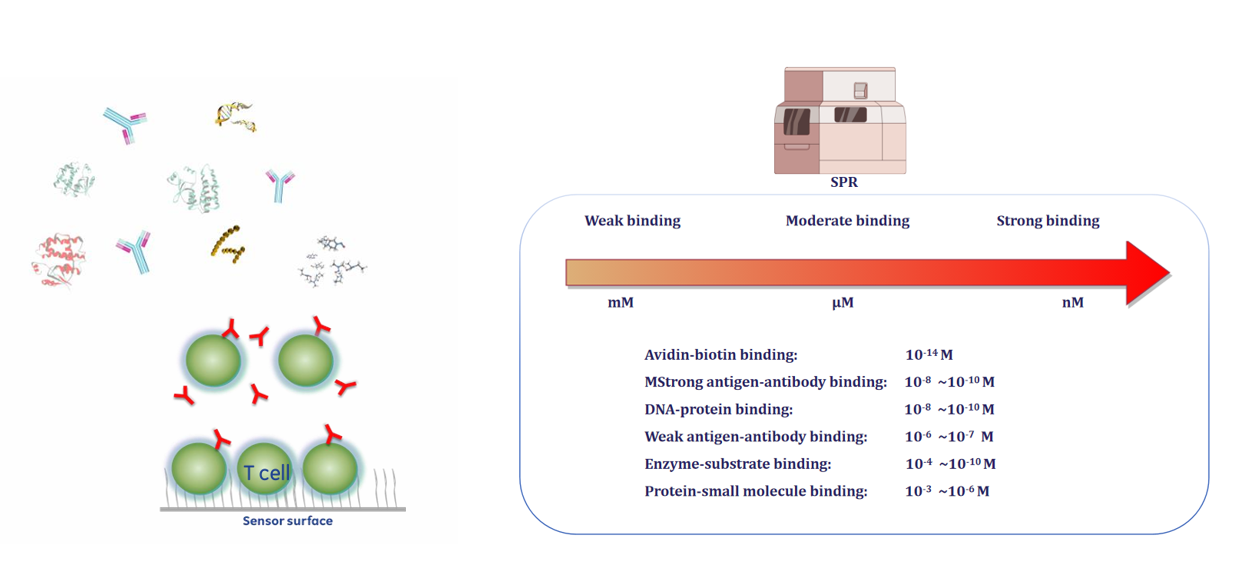
Case:
Protein-Protein Interactions:
a)PD-1 & PD-L1:
Tess Puopolo et al. developed a novel PD-1/PD-L1 blockade assay based on surface plasmon resonance (SPR) technology. This method immobilizes human PD-1 on a sensor chip, allowing it to interact with PD-L1 protein, small molecule inhibitors, or negative controls, and then measures binding kinetics and blockade rates. This protocol is suitable for screening compounds that block the PD-1/PD-L1 interaction. The blocking efficacy of known inhibitors BMS-1166 and BMS-202, as well as the negative control NO-Losartan A, was successfully validated. IC50 values were calculated by calculating the blockade rates. This method has broad application in the discovery of small molecule PD-1/PD-L1 inhibitors for cancer immunotherapy.

(Data source: Tess Puopolo , et al. Bio Protoc. 2023)
b)Entamoeba histolytica EhP3 & Actin regulatory protein Coactosin:
Rath PP et al. used SPR to elucidate the connection between EhP3 and the actin cytoskeleton in the protist pathogen Entamoeba histolytica. They successfully screened several actin-binding/actin-regulating proteins (coactosin, actophorin, twinfilin, profilin, and filamin) that bind to EhP3. Their results revealed coactosin as a key interaction partner of EhP3. A complete kinetic analysis confirmed that the affinity constant of EhCoactosin for EhP3 was 3 μM.
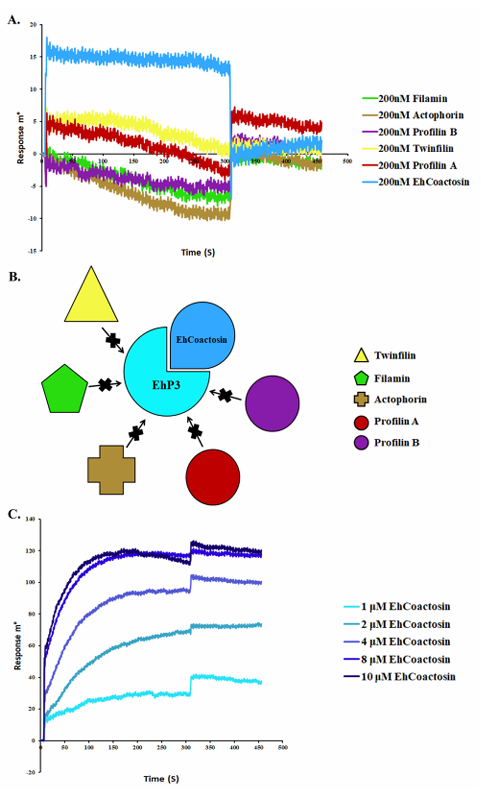
(Data source: Rath PP, et al. Bio Protoc. 2020)
c)Ebola virus glycoprotein GPcl & human cholesterol transporter NPC1-C:
In 2016, a team led by Academician Gao Fu from the Institute of Microbiology, Chinese Academy of Sciences, resolved the 2.3Å resolution crystal structure of EBOV's GPcl bound to the NPC1 - C domain. They discovered that NPC1 - C binds to the hydrophobic cavity of the GPcl head via two protruding loops. Enzymatic cleavage and NPC1 binding trigger conformational changes in GPcl , activating an internal fusion loop and triggering membrane fusion. This revealed the structural mechanism of late endosome entry by filoviruses. Furthermore, the virus first binds to the cell surface (not true entry receptors) via attachment factors such as C-type lectins (e.g. , DC-SIGN and L - SIGN), followed by macropinocytosis-like internalization and early and late endosome transport. This study laid the molecular and structural foundation for the design of therapeutic inhibitors to block viral entry.
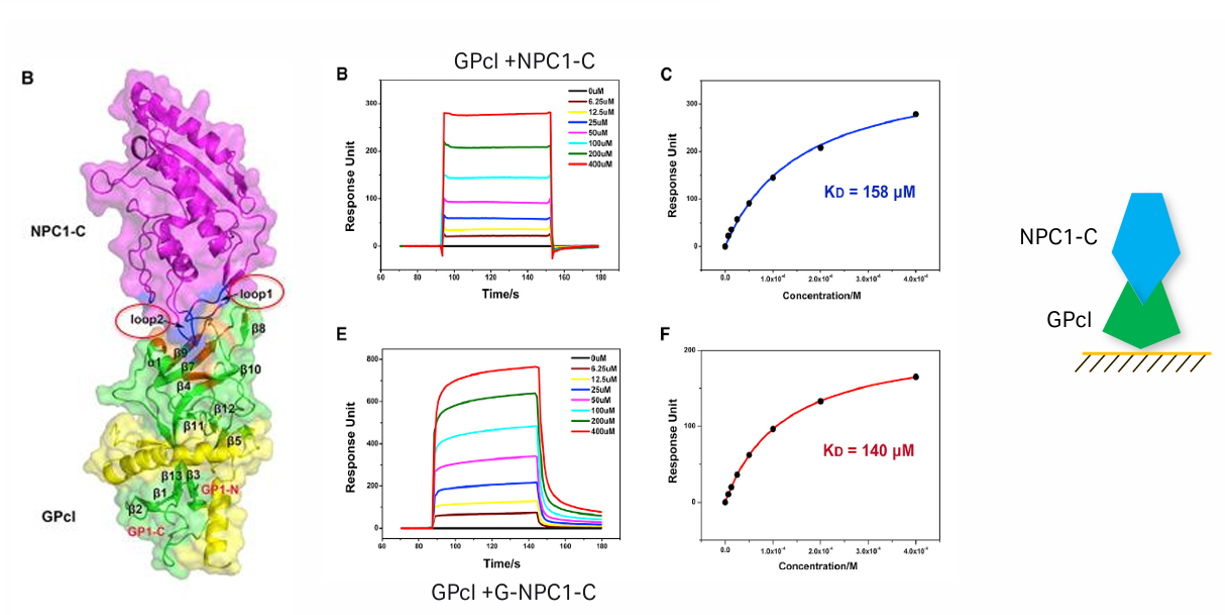
(Data source: Wang H , et al. Cell . 2016)
Protein- nucleic acid interactions:
a)OsPAL6, OsPAL8 gene promoter and transcription factor OsMYB30
The brown planthopper (BPH) is one of the most destructive pests affecting rice (Oryza sativa L.) production. Phenylalanine ammonia-lyase (PAL) is a key enzyme in plant defense against pathogens, but its role in plant resistance remains unclear. In 2019, a team led by Academicians Yuan Longping and Wan Jianmin discovered that brown planthopper feeding significantly induces the expression of most PAL genes (OsPALs) in rice. Silencing OsPALs significantly reduces rice resistance to brown planthoppers, while overexpressing OsPAL8 in susceptible rice varieties significantly enhances resistance. The study demonstrated that the expression levels of phenylalanine ammonia-lyase (PAL)-related genes (OsPAL6 and OsPAL8) are directly upregulated by the rice R2R3 MYB transcription factor OsMYB30. The final research results confirmed that: "The phenylpropanoid metabolic pathway plays an important role in rice's resistance response to brown planthoppers, and at the same time provides valuable target genes for the genetic improvement of rice's brown planthopper resistance."
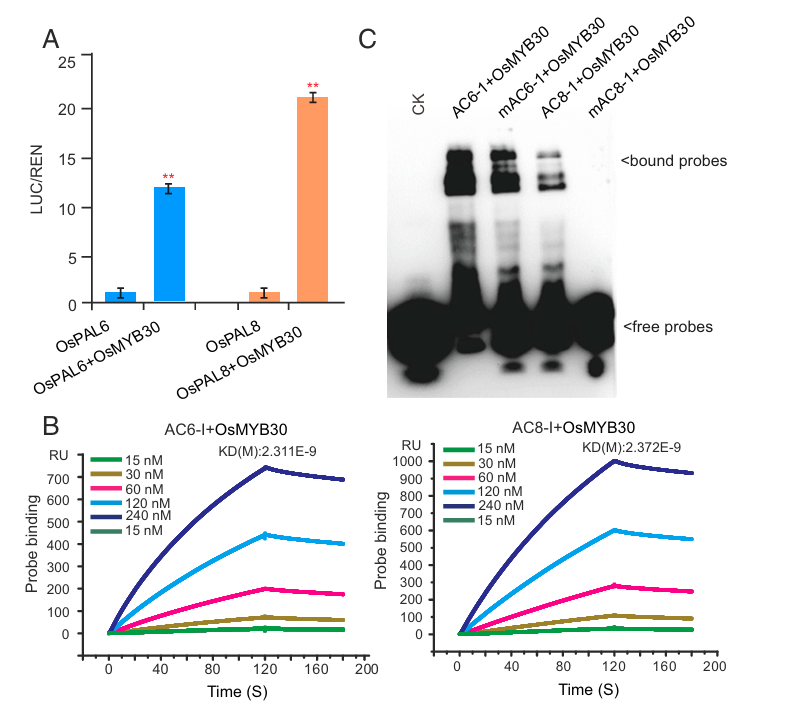
(Data source: He J , et al. Proc Natl Acad Sci USA . 2019)
Protein-polysaccharide/lipid interactions:
a)Capsular polysaccharide CPS & acetylated capsular esterase
Wang Zhe and colleagues used SPR technology to investigate the interaction between Klebsiella pneumoniae capsular polysaccharide (CPS) and acetylated capsular esterase (KpACE). This SPR platform offers short experimental times, minimal sample consumption, and the ability to mimic physiological environments, making it highly suitable for studying carbohydrate-protein interactions. They successfully demonstrated that KpACE targets the acetyl group of CPS, revealing its enzymatic activity and catalytic site, providing key insights into bacterial pathogenicity and antimicrobial drug development. This highly sensitive, label-free method is widely applicable for rapid screening of interaction affinities and enzyme substrate studies.
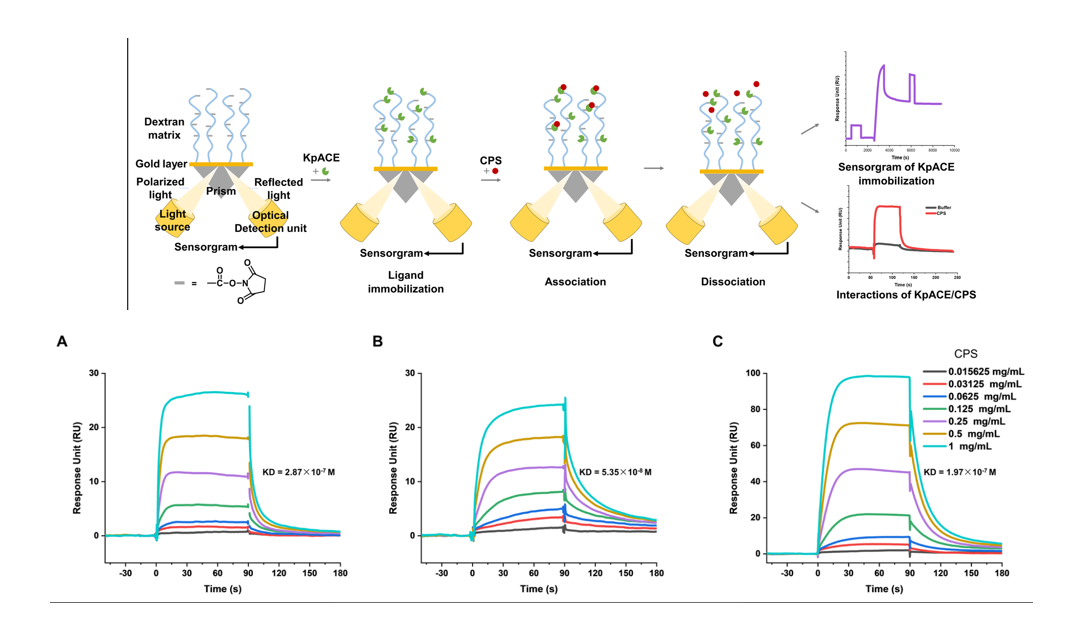
(Data source: Wang Z , et al. Bio Protoc. 2025)
b)Antimicrobial proteins & lipopolysaccharide LPS
The interaction between lipids and proteins is a key event in biological membrane systems. Assessing the affinity and specificity of protein-lipid binding can provide valuable information for elucidating cell membrane function. Baron O'Let et al. described the binding of an antimicrobial protein (ligand) to lipopolysaccharide (LPS) (analyte) after immobilization to a carboxymethyl (CM) sensor chip via amine coupling.
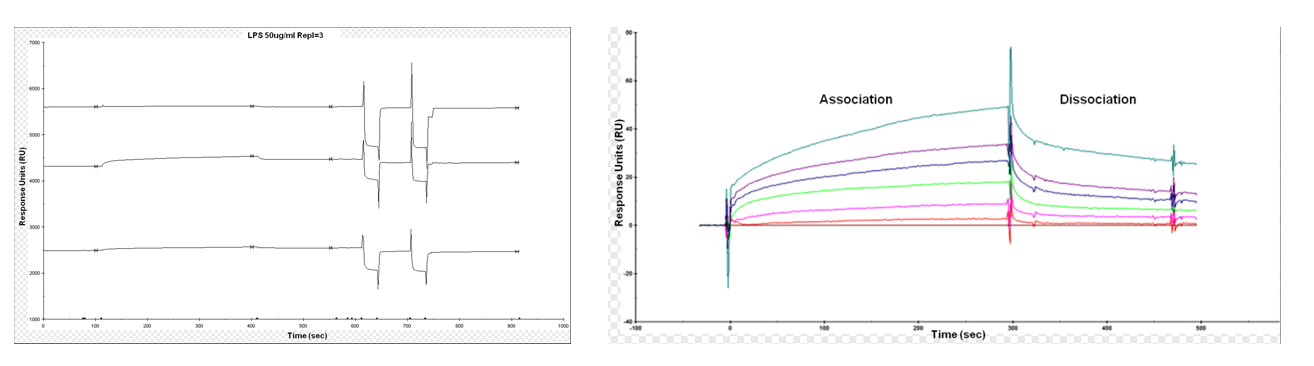
(Data source: Baron OL, et al. Bio Protoc. 2014)
Protein-peptide interactions:
Eiji Ishii et al. successfully established an SPR-based protein-peptide interaction detection method, including ligand immobilization, analyte binding determination, and data analysis steps, which is suitable for quantitatively studying the affinity and kinetic parameters between peptides and proteins.

(Data source: Eiji Ishii , et al. Bio Protoc. 2013)
Protein-Small Molecule Interactions:
a)Small molecule drugs & E3 ligase adaptor protein SPOP:
In 2016, Guo Zhongqiang and his colleagues at the Shanghai Institute of Materia Medica, Chinese Academy of Sciences, successfully designed and optimized a small molecule inhibitor to address the problem of SPOP protein overexpression in the cytoplasm, rather than the nucleus, promoting tumor growth in clear cell renal cell carcinoma (ccRCC). The inhibitor's core mechanism is to block the oncogenic signaling pathway by inhibiting the interaction between SPOP and its substrate proteins. Results showed that the inhibitor effectively killed cancer cells with minimal effects on normal cells, providing a potential new treatment for this cancer.
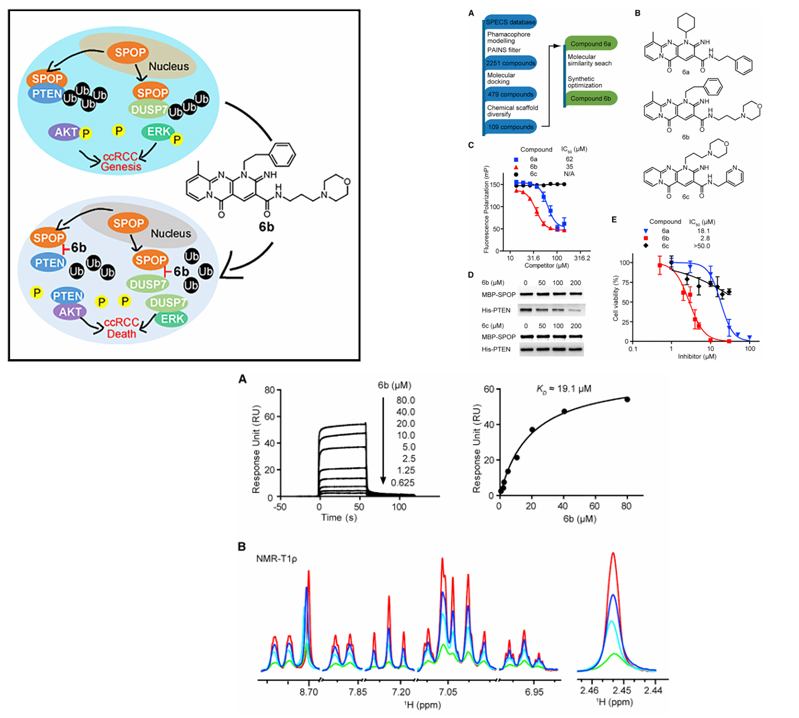
(Data source: Guo ZQ, et al. Cancer Cell. 2016)
b)Triterpenoids &virus fusion protein:
In 2018, Si Longlong's team at Peking University reported a class of natural triterpenoids that target the conserved heptad repeat 2 (HR2), a dominant α-helix structure, in class I viral fusion proteins, thereby antagonizing the formation of hairpin trimers. These compounds inhibit the entry of multiple viruses, including Ebola, Marburg, HIV, and influenza A, with their activity correlated with HR2 sequence characteristics. Using amino acid substitution, surface plasmon resonance, and nuclear magnetic resonance, the study identified six key residues, revealing the molecular mechanism by which the triterpenoids inhibit hairpin formation by binding to the HR2 hydrophobic helix and blocking HR2-HR2 interactions. This mechanism has been validated against both HIV and influenza viruses, suggesting that these three classes of compounds may serve as broad-spectrum antiviral strategies and provide new insights into the regulation of viral fusion mechanisms.
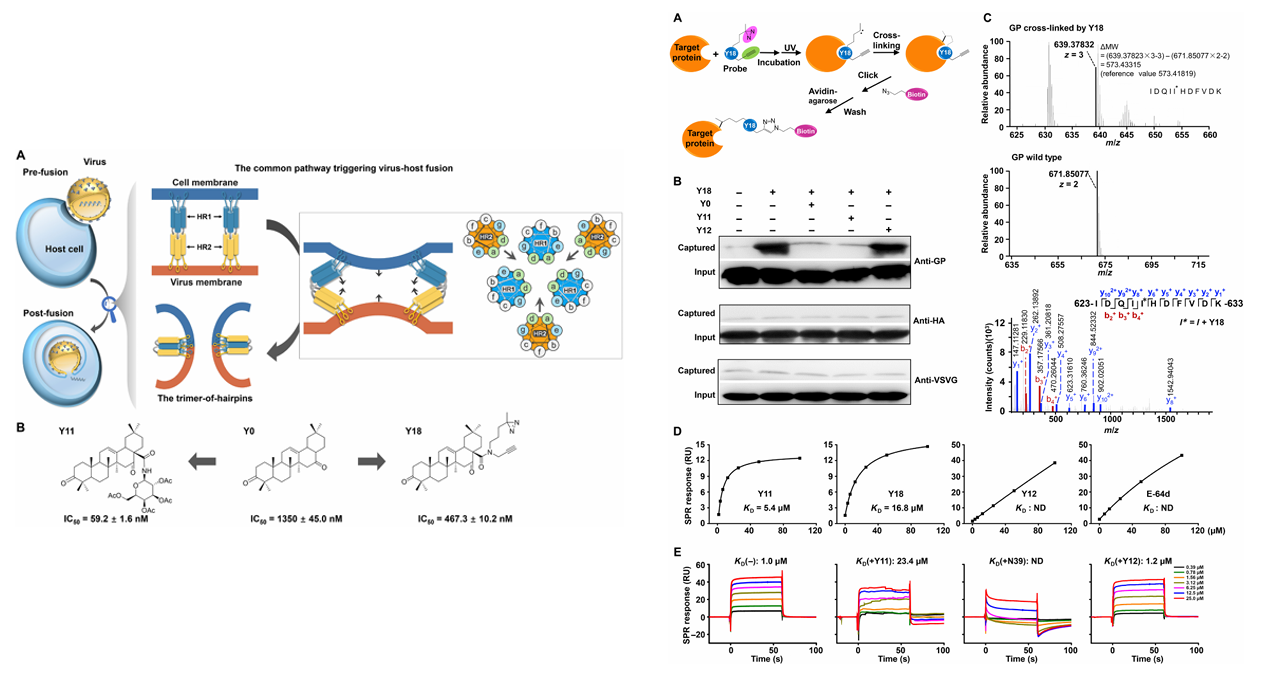
(Data source: Si L , et al. Sci Adv . 2018)
Antigen-antibody interaction:
a)Antigen & Antibody
Nguyen K et al. developed a method for analyzing 1:2 binding kinetics data using a system of ordinary differential equations. Key features of this method include a grid search for parameter initialization and a profile likelihood method for determining parameter identifiability. Using this method, they identified an unidentifiable parameter in a dataset acquired using a standard experimental design. The optimized experimental design, guided by simulations, resulted in reliable estimates of all rate constants.
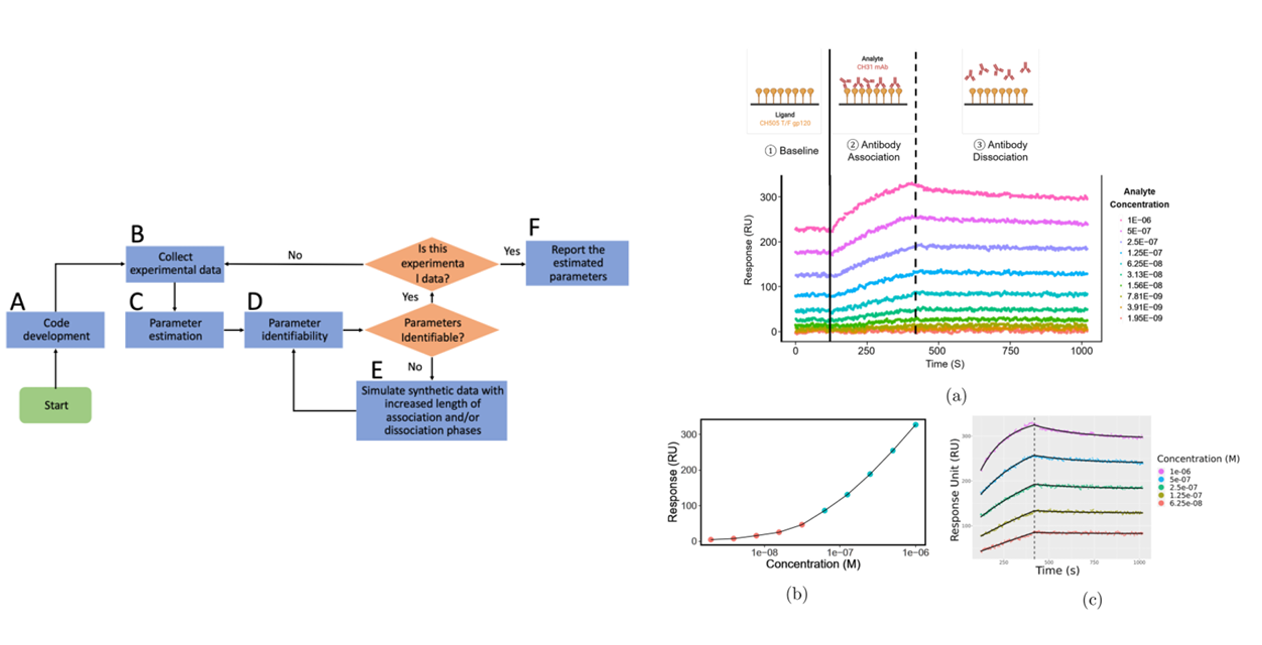
(Data source : Nguyen K, et al. Anal Biochem. 2023)
b)Hybridoma samples & antigens:
Dorin MN et al. used the Biacore T100 system to analyze the affinity of crude hybridoma samples for proteins using SPR technology. This method first covalently immobilized an anti-IgG antibody to a CM5 chip via amide coupling. The Fc region was then used to non-covalently capture the antibody in the hybridoma supernatant, achieving directional binding. To reduce baseline drift, EDC/NHS crosslinking was used to stabilize the complex. KD values were obtained by monitoring the interaction at multiple concentrations and performing kinetic analysis to assess affinity. This strategy avoids the affinity interaction of bivalent antibodies, allows for direct analysis of unpurified samples, and achieves stable, specific antibody capture.
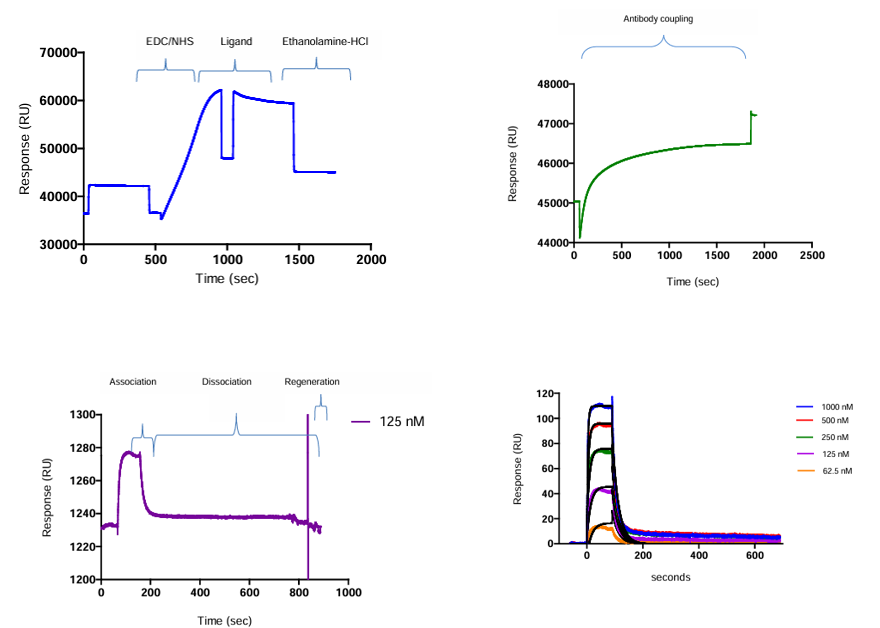
(Data source: Dorin MN, et al. Bio Protoc. 2014)
c)SARS-CoV-2 & Neutralizing Antibodies:
The coronavirus disease 2019 (COVID-19) outbreak, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has spread globally. Countermeasures are needed to treat the disease and prevent further spread of the virus. In 2020, Shi Rui's team at the Institute of Microbiology, Chinese Academy of Sciences, isolated two highly potent neutralizing monoclonal antibodies, CA1 and CB6, from recovered patients. Studies have shown that CB6 significantly inhibits SARS-CoV-2 infection both in vitro and in rhesus macaque models. Structural analysis suggests that CB competitively blocks the virus-receptor interaction by targeting an epitope on the viral RBD that overlaps with the ACE2 binding site. These results highlight the potential of CB6 as a clinical drug candidate, worthy of further development.
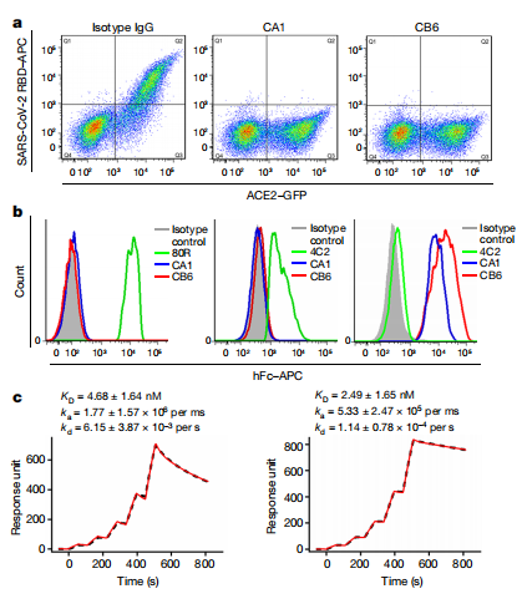
(Data source: Shi R, et al. Nature. 2020)
